A disease-causing mutation in the active site of serine palmitoyltransferase causes catalytic promiscuity
- PMID: 20504773
- PMCID: PMC2906276
- DOI: 10.1074/jbc.M110.122259
A disease-causing mutation in the active site of serine palmitoyltransferase causes catalytic promiscuity
Abstract
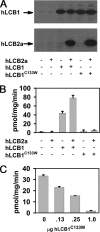
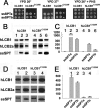
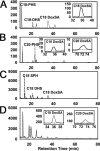
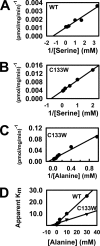
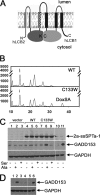
The autosomal dominant peripheral sensory neuropathy HSAN1 results from mutations in the LCB1 subunit of serine palmitoyltransferase (SPT). Serum from patients and transgenic mice expressing a disease-causing mutation (C133W) contain elevated levels of 1-deoxysphinganine (1-deoxySa), which presumably arise from inappropriate condensation of alanine with palmitoyl-CoA. Mutant heterodimeric SPT is catalytically inactive. However, mutant heterotrimeric SPT has approximately 10-20% of wild-type activity and supports growth of yeast cells lacking endogenous SPT. In addition, long chain base profiling revealed the synthesis of significantly more 1-deoxySa in yeast and mammalian cells expressing the heterotrimeric mutant enzyme than in cells expressing wild-type enzyme. Wild-type and mutant enzymes had similar affinities for serine. Surprisingly, the enzymes also had similar affinities for alanine, indicating that the major affect of the C133W mutation is to enhance activation of alanine for condensation with the acyl-CoA substrate. In vivo synthesis of 1-deoxySa by the mutant enzyme was proportional to the ratio of alanine to serine in the growth media, suggesting that this ratio can be used to modulate the relative synthesis of sphinganine and 1-deoxySa. By expressing SPT as a single-chain fusion protein to ensure stoichiometric expression of all three subunits, we showed that GADD153, a marker for endoplasmic reticulum stress, was significantly elevated in cells expressing mutant heterotrimers. GADD153 was also elevated in cells treated with 1-deoxySa. Taken together, these data indicate that the HSAN1 mutations perturb the active site of SPT resulting in a gain of function that is responsible for the HSAN1 phenotype.
Figures

Similar articles
-
HSAN1 mutations in serine palmitoyltransferase reveal a close structure-function-phenotype relationship.Hum Mol Genet. 2016 Mar 1;25(5):853-65. doi: 10.1093/hmg/ddv611. Epub 2015 Dec 17. Hum Mol Genet. 2016. PMID: 26681808
-
The external aldimine form of serine palmitoyltransferase: structural, kinetic, and spectroscopic analysis of the wild-type enzyme and HSAN1 mutant mimics.J Biol Chem. 2009 Jun 19;284(25):17328-17339. doi: 10.1074/jbc.M109.008680. Epub 2009 Apr 17. J Biol Chem. 2009. PMID: 19376777 Free PMC article.
-
Mutant SPTLC1 dominantly inhibits serine palmitoyltransferase activity in vivo and confers an age-dependent neuropathy.Hum Mol Genet. 2005 Nov 15;14(22):3507-21. doi: 10.1093/hmg/ddi380. Epub 2005 Oct 6. Hum Mol Genet. 2005. PMID: 16210380
-
Structural, mechanistic and regulatory studies of serine palmitoyltransferase.Biochem Soc Trans. 2012 Jun 1;40(3):547-54. doi: 10.1042/BST20110769. Biochem Soc Trans. 2012. PMID: 22616865 Review.
-
Serine palmitoyltransferase, a key enzyme of sphingolipid metabolism.Biochim Biophys Acta. 2003 Jun 10;1632(1-3):16-30. doi: 10.1016/s1388-1981(03)00059-3. Biochim Biophys Acta. 2003. PMID: 12782147 Review.
Cited by
-
ORM Expression Alters Sphingolipid Homeostasis and Differentially Affects Ceramide Synthase Activity.Plant Physiol. 2016 Oct;172(2):889-900. doi: 10.1104/pp.16.00965. Epub 2016 Aug 9. Plant Physiol. 2016. PMID: 27506241 Free PMC article.
-
Druggable Sphingolipid Pathways: Experimental Models and Clinical Opportunities.Adv Exp Med Biol. 2020;1274:101-135. doi: 10.1007/978-3-030-50621-6_6. Adv Exp Med Biol. 2020. PMID: 32894509 Review.
-
Quantifying 1-deoxydihydroceramides and 1-deoxyceramides in mouse nervous system tissue.Prostaglandins Other Lipid Mediat. 2019 Apr;141:40-48. doi: 10.1016/j.prostaglandins.2019.02.005. Epub 2019 Feb 19. Prostaglandins Other Lipid Mediat. 2019. PMID: 30790665 Free PMC article.
-
Autophagy regulates sphingolipid levels in the liver.J Lipid Res. 2014 Dec;55(12):2521-31. doi: 10.1194/jlr.M051862. Epub 2014 Oct 20. J Lipid Res. 2014. PMID: 25332431 Free PMC article.
-
Allosteric conformational barcodes direct signaling in the cell.Structure. 2013 Sep 3;21(9):1509-21. doi: 10.1016/j.str.2013.06.002. Structure. 2013. PMID: 24010710 Free PMC article. Review.
References
-
- Hornemann T., Richard S., Rütti M. F., Wei Y., von Eckardstein A. (2006) J. Biol. Chem. 281, 37275–37281 - PubMed
-
- Bejaoui K., Wu C., Scheffler M. D., Haan G., Ashby P., Wu L., de Jong P., Brown R. H., Jr. (2001) Nat. Genet. 27, 261–262 - PubMed
-
- Dawkins J. L., Hulme D. J., Brahmbhatt S. B., Auer-Grumbach M., Nicholson G. A. (2001) Nat. Genet. 27, 309–312 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

