Use of dimedone-based chemical probes for sulfenic acid detection methods to visualize and identify labeled proteins
- PMID: 20513473
- PMCID: PMC3835715
- DOI: 10.1016/S0076-6879(10)73004-4
Use of dimedone-based chemical probes for sulfenic acid detection methods to visualize and identify labeled proteins
Abstract
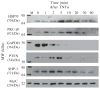
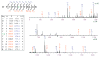
Reversible thiol modification is a major component of the modulation of cell-signaling pathways by reactive oxygen species. Hydrogen peroxide, peroxynitrite, or lipid hydroperoxides are all able to oxidize cysteines to form cysteine sulfenic acids; this reactive intermediate can be directly reduced to thiol by cellular reductants such as thioredoxin or further participate in disulfide bond formation with glutathione or cysteine residues in the same or another protein. To identify the direct protein targets of cysteine modification and the conditions under which they are oxidized, a series of dimedone-based reagents linked to affinity or fluorescent tags have been developed that specifically alkylate and trap cysteine sulfenic acids. In this chapter, we provide detailed methods using one of our biotin-tagged reagents, DCP-Bio1, to identify and monitor proteins that are oxidized in vitro and in vivo. Using streptavidin-linked agarose beads, this biotin-linked reagent can be used to affinity capture labeled proteins. Stringent washing of the beads prior to elution minimizes the contamination of the enriched material with unlabeled proteins through coimmunoprecipitation or nonspecific binding. In particular, we suggest including DTT in one of the washes to remove proteins covalently linked to biotinylated proteins through a disulfide bond, except in cases where these linked proteins are of interest. We also provide methods for targeted approaches monitoring cysteine oxidation in individual proteins, global approaches to follow total cysteine oxidation in the cell, and guidelines for proteomic analyses to identify novel proteins with redox sensitive cysteines.
Copyright (c) 2010 Elsevier Inc. All rights reserved.
Figures
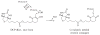



References
-
- Chen CH, Cheng TH, Lin H, Shih NL, Chen YL, Chen YS, Cheng CF, Lian WS, Meng TC, Chiu WT, Chen JJ. Reactive oxygen species generation is involved in epidermal growth factor receptor transactivation through the transient oxidization of Src homology 2-containing tyrosine phosphatase in endothelin-1 signaling pathway in rat cardiac fibroblasts. Mol Pharmacol. 2006;69:1347–1355. - PubMed
-
- Claiborne A, Yeh JI, Mallett TC, Luba J, Crane EJ, 3rd, Charrier V, Parsonage D. Protein-sulfenic acids: Diverse roles for an unlikely player in enzyme catalysis and redox regulation. Biochemistry. 1999;38:15407–15416. - PubMed
-
- Cotgreave IA, Gerdes R, Schuppe-Koistinen I, Lind C. S-glutathionylation of glyceraldehyde-3-phosphate dehydrogenase: Role of thiol oxidation and catalysis by glutaredoxin. Methods Enzymol. 2002;348:175–182. - PubMed
-
- Dennehy MK, Richards KA, Wernke GR, Shyr Y, Liebler DC. Cytosolic and nuclear protein targets of thiol-reactive electrophiles. Chem Res Toxicol. 2006;19:20–29. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

