Using environmental correlations to identify loci underlying local adaptation
- PMID: 20516501
- PMCID: PMC2927766
- DOI: 10.1534/genetics.110.114819
Using environmental correlations to identify loci underlying local adaptation
Abstract
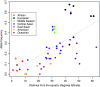
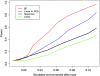
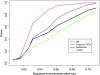
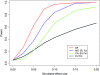
Loci involved in local adaptation can potentially be identified by an unusual correlation between allele frequencies and important ecological variables or by extreme allele frequency differences between geographic regions. However, such comparisons are complicated by differences in sample sizes and the neutral correlation of allele frequencies across populations due to shared history and gene flow. To overcome these difficulties, we have developed a Bayesian method that estimates the empirical pattern of covariance in allele frequencies between populations from a set of markers and then uses this as a null model for a test at individual SNPs. In our model the sample frequencies of an allele across populations are drawn from a set of underlying population frequencies; a transform of these population frequencies is assumed to follow a multivariate normal distribution. We first estimate the covariance matrix of this multivariate normal across loci using a Monte Carlo Markov chain. At each SNP, we then provide a measure of the support, a Bayes factor, for a model where an environmental variable has a linear effect on the transformed allele frequencies compared to a model given by the covariance matrix alone. This test is shown through power simulations to outperform existing correlation tests. We also demonstrate that our method can be used to identify SNPs with unusually large allele frequency differentiation and offers a powerful alternative to tests based on pairwise or global F(ST). Software is available at http://www.eve.ucdavis.edu/gmcoop/.
Figures

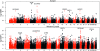
References
-
- Allen, J., 1877. The influence of physical conditions in the genesis of species. Radic. Rev. 1 108–140.
-
- Balding, D., 2003. Likelihood-based inference for genetic correlation coefficients. Theor. Popul. Biol. 63 221–230. - PubMed
-
- Balding, D. J., and R. A. Nichols, 1995. A method for quantifying differentiation between populations at multi-allelic loci and its implications for investigating identity and paternity. Genetica 96 3–12. - PubMed
-
- Balding, D. J., A. D. Carothers, J. L. Marchini, L. R. Cardon, A. Vetta et al., 2002. Discussion on the meeting on ‘Statistical modelling and analysis of genetic data’. J. R. Stat. Soc. B 64 737–775.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

