The neurogenic basic helix-loop-helix transcription factor NeuroD6 confers tolerance to oxidative stress by triggering an antioxidant response and sustaining the mitochondrial biomass
- PMID: 20517466
- PMCID: PMC2874871
- DOI: 10.1042/AN20100005
The neurogenic basic helix-loop-helix transcription factor NeuroD6 confers tolerance to oxidative stress by triggering an antioxidant response and sustaining the mitochondrial biomass
Abstract
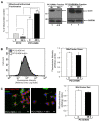
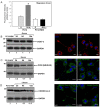
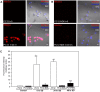
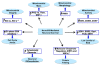
Preserving mitochondrial mass, bioenergetic functions and ROS (reactive oxygen species) homoeostasis is key to neuronal differentiation and survival, as mitochondria produce most of the energy in the form of ATP to execute and maintain these cellular processes. In view of our previous studies showing that NeuroD6 promotes neuronal differentiation and survival on trophic factor withdrawal, combined with its ability to stimulate the mitochondrial biomass and to trigger comprehensive antiapoptotic and molecular chaperone responses, we investigated whether NeuroD6 could concomitantly modulate the mitochondrial biomass and ROS homoeostasis on oxidative stress mediated by serum deprivation. In the present study, we report a novel role of NeuroD6 as a regulator of ROS homoeostasis, resulting in enhanced tolerance to oxidative stress. Using a combination of flow cytometry, confocal fluorescence microscopy and mitochondrial fractionation, we found that NeuroD6 sustains mitochondrial mass, intracellular ATP levels and expression of specific subunits of respiratory complexes upon oxidative stress triggered by withdrawal of trophic factors. NeuroD6 also maintains the expression of nuclear-encoded transcription factors, known to regulate mitochondrial biogenesis, such as PGC-1alpha (peroxisome-proliferator-activated receptor gamma co-activator-1alpha), Tfam (transcription factor A, mitochondrial) and NRF-1 (nuclear respiratory factor-1). Finally, NeuroD6 triggers a comprehensive antioxidant response to endow PC12-ND6 cells with intracellular ROS scavenging capacity. The NeuroD6 effect is not limited to the classic induction of the ROS-scavenging enzymes, such as SOD2 (superoxide dismutase 2), GPx1 (glutathione peroxidase 1) and PRDX5 (peroxiredoxin 5), but also to the recently identified powerful ROS suppressors PGC-1alpha, PINK1 (phosphatase and tensin homologue-induced kinase 1) and SIRT1. Thus our collective results support the concept that the NeuroD6-PGC-1alpha-SIRT1 neuroprotective axis may be critical in co-ordinating the mitochondrial biomass with the antioxidant reserve to confer tolerance to oxidative stress.
Keywords: AD, Alzheimer’s disease; AM, acetoxymethyl ester; COX, cytochrome c oxidase; DAPI, 4′,6-diamidino-2-phenylindole; DIC, differential interference contrast; Drp1, dynamin-related protein 1; ETC, electron transfer chain; GABP-α, GA-binding protein-α; GAPDH, glyceraldehyde-3-phosphate dehydrogenase; GFP, green fluorescent protein; GPx1, glutathione peroxidase 1; HSP, heat-shock protein; MMP, mitochondrial membrane potential; MTG, MitoTracker® Green; MTR, MitoTracker® Red; Mfn2, mitofusin 2; Mg-Gr, Magnesium Green; NRF, nuclear respiratory factor; NT-PGC-1α, N-terminal-truncated PGC-1α; NeuroD family; OPA1, optic atrophy 1; OXPHOS, oxidative phosphorylation; PDL, poly-d-lysine; PGC-1α, peroxisome-proliferator-activated receptor γ co-activator-1α; PINK1, phosphatase and tensin homologue-induced kinase 1; PRDX5, peroxiredoxin 5; ROS, reactive oxygen species; SIRT1; SOD, superoxide dismutase; Tfam, transcription factor A, mitochondrial; WGA, wheatgerm agglutinin; bHLH, basic helix–loop–helix; mitochondria; mtDNA, mitochondrial DNA; neuronal survival; reactive oxygen species (ROS); transcriptional co-regulator peroxisome-proliferator-activated receptor γ co-activator-1α (PGC-1α).
Figures



Similar articles
-
The neurogenic basic helix-loop-helix transcription factor NeuroD6 concomitantly increases mitochondrial mass and regulates cytoskeletal organization in the early stages of neuronal differentiation.ASN Neuro. 2009 Sep 16;1(4):e00016. doi: 10.1042/AN20090036. ASN Neuro. 2009. PMID: 19743964 Free PMC article.
-
The neurogenic basic helix-loop-helix transcription factor NeuroD6 enhances mitochondrial biogenesis and bioenergetics to confer tolerance of neuronal PC12-NeuroD6 cells to the mitochondrial stressor rotenone.Exp Cell Res. 2012 Oct 15;318(17):2200-14. doi: 10.1016/j.yexcr.2012.07.004. Epub 2012 Jul 16. Exp Cell Res. 2012. PMID: 22814253 Free PMC article.
-
Mitochondrial biogenesis: pharmacological approaches.Curr Pharm Des. 2014;20(35):5507-9. doi: 10.2174/138161282035140911142118. Curr Pharm Des. 2014. PMID: 24606795
-
Natural products, PGC-1 α , and Duchenne muscular dystrophy.Acta Pharm Sin B. 2020 May;10(5):734-745. doi: 10.1016/j.apsb.2020.01.001. Epub 2020 Jan 8. Acta Pharm Sin B. 2020. PMID: 32528825 Free PMC article. Review.
-
Roles of oxidative stress, apoptosis, PGC-1α and mitochondrial biogenesis in cerebral ischemia.Int J Mol Sci. 2011;12(10):7199-215. doi: 10.3390/ijms12107199. Epub 2011 Oct 21. Int J Mol Sci. 2011. PMID: 22072942 Free PMC article. Review.
Cited by
-
The emerging roles of microRNAs in CNS injuries.Nat Rev Neurol. 2013 Jun;9(6):328-39. doi: 10.1038/nrneurol.2013.67. Epub 2013 Apr 16. Nat Rev Neurol. 2013. PMID: 23588363 Free PMC article. Review.
-
Indomethacin promotes survival of new neurons in the adult murine hippocampus accompanied by anti-inflammatory effects following MPTP-induced dopamine depletion.J Neuroinflammation. 2018 May 26;15(1):162. doi: 10.1186/s12974-018-1179-4. J Neuroinflammation. 2018. PMID: 29803225 Free PMC article.
-
microRNAs in spinal cord injury: potential roles and therapeutic implications.Int J Biol Sci. 2014 Sep 6;10(9):997-1006. doi: 10.7150/ijbs.9058. eCollection 2014. Int J Biol Sci. 2014. PMID: 25210498 Free PMC article. Review.
-
Generation of a DAT-P2A-Flpo mouse line for intersectional genetic targeting of dopamine neuron subpopulations.Cell Rep. 2021 May 11;35(6):109123. doi: 10.1016/j.celrep.2021.109123. Cell Rep. 2021. PMID: 33979604 Free PMC article.
-
Role of Cell Metabolism and Mitochondrial Function During Adult Neurogenesis.Neurochem Res. 2017 Jun;42(6):1787-1794. doi: 10.1007/s11064-016-2150-3. Epub 2016 Dec 21. Neurochem Res. 2017. PMID: 28000162 Review.
References
-
- Balaban RS, Nemoto S, Finkel T. Mitochondria, oxidants, and aging. Cell. 2005;120:483–495. - PubMed
-
- Banmeyer I, Marchand C, Clippe A, Knoops B. Human mitochondrial peroxiredoxin 5 protects from mitochondrial DNA damages induced by hydrogen peroxide. FEBS Lett. 2005;579:2327–2333. - PubMed
-
- Banmeyer I, Marchand C, Verhaeghe C, Vucic B, Rees JF, Knoops B. Overexpression of human peroxiredoxin 5 in subcellular compartments of chinese hamster ovary cells: Effects on cytotoxicity and DNA damage caused by peroxides. Free Radical Biol Med. 2004;36:65–77. - PubMed
-
- Bartholomä A, Nave KA. NEX-1: a novel brain-specific helix–loop–helix protein with autoregulation and sustained expression in mature cortical neurons. Mech Dev. 1994;48:217–228. - PubMed
-
- Batchelor AH, Piper DE, de la Brousse FC, McKnight SL, Wolberger C. The structure of GABPalpha/beta: an ETS domain–ankyrin repeat heterodimer bound to DNA. Science. 1998;279:1037–1041. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

