Using functional genetics to understand breast cancer biology
- PMID: 20519343
- PMCID: PMC2890198
- DOI: 10.1101/cshperspect.a003327
Using functional genetics to understand breast cancer biology
Abstract
Genetic screens were for long the prerogative of those that studied model organisms. The discovery in 2001 that gene silencing through RNA interference (RNAi) can also be brought about in mammalian cells paved the way for large scale loss-of-function genetic screens in higher organisms. In this article, we describe how functional genetic studies can help us understand the biology of breast cancer, how it can be used to identify novel targets for breast cancer therapy, and how it can help in the identification of those patients that are most likely to respond to a given therapy.
Figures

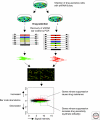

References
-
- Bernards R, Brummelkamp TR, Beijersbergen RL 2006. shRNA libraries and their use in cancer genetics. Nat Methods 3:701–706 - PubMed
-
- Berns K, Hijmans EM, Mullenders J, Brummelkamp TR, Velds A, Heimerikx M, Kerkhoven RM, Madiredjo M, Nijkamp W, Weigelt B, et al.2004. A large-scale RNAi screen in human cells identifies new components of the p53 pathway. Nature 428:431–437 - PubMed
-
- Berns K, Horlings HM, Hennessy BT, Madiredjo M, Hijmans EM, Beelen K, Linn SC, Gonzalez-Angulo AM, Stemke-Hale K, Hauptmann M, et al.2007. A functional genetic approach identifies the PI3K pathway as a major determinant of trastuzumab resistance in breast cancer. Cancer Cell 12:395–402 - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
