Ontologies in quantitative biology: a basis for comparison, integration, and discovery
- PMID: 20520843
- PMCID: PMC2876043
- DOI: 10.1371/journal.pbio.1000374
Ontologies in quantitative biology: a basis for comparison, integration, and discovery
Abstract
As biology is becoming a data-driven discipline, ontologies become increasingly important for systematically capturing the existing knowledge. This essay discusses current trends and how ontologies can also be used for discovery.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
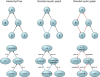

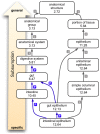
References
-
- Gruber T. R. A translation approach to portable ontology specifications. Knowledge Acquisition. 1993;5:199–220.
-
- Taylor W. R. The classification of amino acid conservation. J Theor Biol. 1986;119:205–218. - PubMed
-
- Murzin A. G, Brenner S. E, Hubbard T, Chothia C. SCOP: a structural classification of proteins database for the investigation of sequences and structures. J Mol Biol. 1995;247:536–540. - PubMed
-
- Orengo C. A, Michie A. D, Jones S, Jones D. T, Swindells M. B, et al. CATH–a hierarchic classification of protein domain structures. Structure. 1997;5:1093–1108. - PubMed
-
- Sonnhammer E. L, Eddy S. R, Durbin R. Pfam: a comprehensive database of protein domain families based on seed alignments. Proteins. 1997;28:405–420. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Research Materials

