A software tool to model genetic regulatory networks. Applications to the modeling of threshold phenomena and of spatial patterning in Drosophila
- PMID: 20523731
- PMCID: PMC2877713
- DOI: 10.1371/journal.pone.0010743
A software tool to model genetic regulatory networks. Applications to the modeling of threshold phenomena and of spatial patterning in Drosophila
Abstract
We present a general methodology in order to build mathematical models of genetic regulatory networks. This approach is based on the mass action law and on the Jacob and Monod operon model. The mathematical models are built symbolically by the Mathematica software package GeneticNetworks. This package accepts as input the interaction graphs of the transcriptional activators and repressors of a biological process and, as output, gives the mathematical model in the form of a system of ordinary differential equations. All the relevant biological parameters are chosen automatically by the software. Within this framework, we show that concentration dependent threshold effects in biology emerge from the catalytic properties of genes and its associated conservation laws. We apply this methodology to the segment patterning in Drosophila early development and we calibrate the genetic transcriptional network responsible for the patterning of the gap gene proteins Hunchback and Knirps, along the antero-posterior axis of the Drosophila embryo. In this approach, the zygotically produced proteins Hunchback and Knirps do not diffuse along the antero-posterior axis of the embryo of Drosophila, developing a spatial pattern due to concentration dependent thresholds. This shows that patterning at the gap genes stage can be explained by the concentration gradients along the embryo of the transcriptional regulators.
Conflict of interest statement
Figures
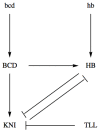
 . Lines with perpendicular endings represent repressions and are listed in the set
. Lines with perpendicular endings represent repressions and are listed in the set  .
.


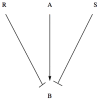
 , the equilibrium value of protein concentration
, the equilibrium value of protein concentration  is zero, while above it takes the value
is zero, while above it takes the value  . The parameters are:
. The parameters are:  ,
,  ,
,  and
and  .
.
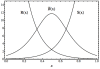
 shows a spiky profile, resulting from the inhibitory action of proteins
shows a spiky profile, resulting from the inhibitory action of proteins  and
and  . In this model, we have considered that the concentrations of
. In this model, we have considered that the concentrations of  and
and  are constant in time and non-homogeneous in space. The activator protein
are constant in time and non-homogeneous in space. The activator protein  has been considered constant along the spatial region.
has been considered constant along the spatial region.
 to
to  , and the units in the vertical axis are proportional to protein concentration. The data has been taken from the FlyEx database. The continuous curves represent the fitted mean distribution of the concentration of proteins calculated from the data of
, and the units in the vertical axis are proportional to protein concentration. The data has been taken from the FlyEx database. The continuous curves represent the fitted mean distribution of the concentration of proteins calculated from the data of  embryos. These curves are the initial conditions for a model obtained with the software package GeneticNetworks for the production of the proteins KNI and zygotically produced HB, during cleavage cycle 14.
embryos. These curves are the initial conditions for a model obtained with the software package GeneticNetworks for the production of the proteins KNI and zygotically produced HB, during cleavage cycle 14.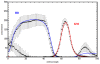
 embryos, taken from the FlyEx database. The embryo length has been scaled from
embryos, taken from the FlyEx database. The embryo length has been scaled from  to
to  , and the units in the vertical axis are proportional to protein concentration. The continuous curves show the predictions of the model of production of proteins HB and KNI. The model equations have been obtained with the software package GeneticNetworks.
, and the units in the vertical axis are proportional to protein concentration. The continuous curves show the predictions of the model of production of proteins HB and KNI. The model equations have been obtained with the software package GeneticNetworks.Similar articles
-
Whole-embryo modeling of early segmentation in Drosophila identifies robust and fragile expression domains.Biophys J. 2011 Jul 20;101(2):287-96. doi: 10.1016/j.bpj.2011.05.060. Biophys J. 2011. PMID: 21767480 Free PMC article.
-
Spatial bistability generates hunchback expression sharpness in the Drosophila embryo.PLoS Comput Biol. 2008 Sep 26;4(9):e1000184. doi: 10.1371/journal.pcbi.1000184. PLoS Comput Biol. 2008. PMID: 18818726 Free PMC article.
-
Boolean modeling of biological regulatory networks: a methodology tutorial.Methods. 2013 Jul 15;62(1):3-12. doi: 10.1016/j.ymeth.2012.10.012. Epub 2012 Nov 9. Methods. 2013. PMID: 23142247
-
The gap gene network.Cell Mol Life Sci. 2011 Jan;68(2):243-74. doi: 10.1007/s00018-010-0536-y. Epub 2010 Oct 8. Cell Mol Life Sci. 2011. PMID: 20927566 Free PMC article. Review.
-
Drosophila blastoderm patterning.Curr Opin Genet Dev. 2012 Dec;22(6):533-41. doi: 10.1016/j.gde.2012.10.005. Epub 2013 Jan 4. Curr Opin Genet Dev. 2012. PMID: 23290311 Review.
Cited by
-
The Drosophila gap gene network is composed of two parallel toggle switches.PLoS One. 2011;6(7):e21145. doi: 10.1371/journal.pone.0021145. Epub 2011 Jul 1. PLoS One. 2011. PMID: 21747931 Free PMC article.
-
Inference of complex biological networks: distinguishability issues and optimization-based solutions.BMC Syst Biol. 2011 Oct 28;5:177. doi: 10.1186/1752-0509-5-177. BMC Syst Biol. 2011. PMID: 22034917 Free PMC article.
-
Quantification reveals early dynamics in Drosophila maternal gradients.PLoS One. 2021 Aug 19;16(8):e0244701. doi: 10.1371/journal.pone.0244701. eCollection 2021. PLoS One. 2021. PMID: 34411119 Free PMC article.
-
Fibroblast state switching orchestrates dermal maturation and wound healing.Mol Syst Biol. 2018 Aug 29;14(8):e8174. doi: 10.15252/msb.20178174. Mol Syst Biol. 2018. PMID: 30158243 Free PMC article.
References
-
- Sánchez L, Thieffry D. A logical analysis of the Drosophila gap-gene system. J Theo Bio. 2001;211:115–141. - PubMed
-
- Alves F, Dilão R. Modeling segmental patterning in Drosophila: Maternal and gap genes. J Theo Bio. 2006;241:342–359. - PubMed
-
- Jong HD. Modelling and simulations of genetic regulatory systems: a literature review. J Comput Biol. 2002;9:67–103. - PubMed
-
- Klipp E, Herwig R, Kowald A, Wierling C, Lehrach H. Systems Biology in Practice. Weinheim: Wiley-VCH; 2005. pp. 282–286.
-
- van Kampen NG. Stochastic Processes in Physics and Chemistry. Amsterdam: North-Holland; 1992. pp. 166–173.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

