Sgs1 and exo1 redundantly inhibit break-induced replication and de novo telomere addition at broken chromosome ends
- PMID: 20523895
- PMCID: PMC2877739
- DOI: 10.1371/journal.pgen.1000973
Sgs1 and exo1 redundantly inhibit break-induced replication and de novo telomere addition at broken chromosome ends
Abstract
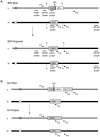
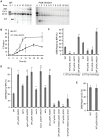
In budding yeast, an HO endonuclease-inducible double-strand break (DSB) is efficiently repaired by several homologous recombination (HR) pathways. In contrast to gene conversion (GC), where both ends of the DSB can recombine with the same template, break-induced replication (BIR) occurs when only the centromere-proximal end of the DSB can locate homologous sequences. Whereas GC results in a small patch of new DNA synthesis, BIR leads to a nonreciprocal translocation. The requirements for completing BIR are significantly different from those of GC, but both processes require 5' to 3' resection of DSB ends to create single-stranded DNA that leads to formation of a Rad51 filament required to initiate HR. Resection proceeds by two pathways dependent on Exo1 or the BLM homolog, Sgs1. We report that Exo1 and Sgs1 each inhibit BIR but have little effect on GC, while overexpression of either protein severely inhibits BIR. In contrast, overexpression of Rad51 markedly increases the efficiency of BIR, again with little effect on GC. In sgs1Delta exo1Delta strains, where there is little 5' to 3' resection, the level of BIR is not different from either single mutant; surprisingly, there is a two-fold increase in cell viability after HO induction whereby 40% of all cells survive by formation of a new telomere within a few kb of the site of DNA cleavage. De novo telomere addition is rare in wild-type, sgs1Delta, or exo1Delta cells. In sgs1Delta exo1Delta, repair by GC is severely inhibited, but cell viability remains high because of new telomere formation. These data suggest that the extensive 5' to 3' resection that occurs before the initiation of new DNA synthesis in BIR may prevent efficient maintenance of a Rad51 filament near the DSB end. The severe constraint on 5' to 3' resection, which also abrogates activation of the Mec1-dependent DNA damage checkpoint, permits an unprecedented level of new telomere addition.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




References
-
- San Filippo J, Sung P, Klein H. Mechanism of eukaryotic homologous recombination. Annu Rev Biochem. 2008;77:229–257. - PubMed
-
- Krogh BO, Symington LS. Recombination proteins in yeast. Annu Rev Genet. 2004;38:233–271. - PubMed
-
- Llorente B, Smith CE, Symington LS. Break-induced replication: what is it and what is it for? Cell Cycle. 2008;7:859–864. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

