Germinal center T follicular helper cell IL-4 production is dependent on signaling lymphocytic activation molecule receptor (CD150)
- PMID: 20525889
- PMCID: PMC2913439
- DOI: 10.4049/jimmunol.0903505
Germinal center T follicular helper cell IL-4 production is dependent on signaling lymphocytic activation molecule receptor (CD150)
Abstract
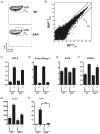
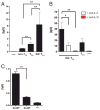
CD4 T cell help is critical for the generation and maintenance of germinal centers (GCs), and T follicular helper (T(FH)) cells are the CD4 T cell subset required for this process. Signaling lymphocytic activation molecule (SLAM)-associated protein (SAP [SH2D1A]) expression in CD4 T cells is essential for GC development. However, SAP-deficient mice have only a moderate defect in T(FH) differentiation, as defined by common T(FH) surface markers. CXCR5(+) T(FH) cells are found within the GC, as well as along the boundary regions of T/B cell zones. In this study, we show that GC-associated T follicular helper (GC T(FH)) cells can be identified by their coexpression of CXCR5 and the GL7 epitope, allowing for phenotypic and functional analysis of T(FH) and GC T(FH) populations. GC T(FH) cells are a functionally discrete subset of further polarized T(FH) cells, with enhanced B cell help capacity and a specialized ability to produce IL-4 in a T(H)2-independent manner. Strikingly, SAP-deficient mice have an absence of the GC T(FH) cell subset and SAP(-) T(FH) cells are defective in IL-4 and IL-21 production. We further demonstrate that SLAM (Slamf1, CD150), a surface receptor that uses SAP signaling, is specifically required for IL-4 production by GC T(FH) cells. GC T(FH) cells require IL-4 and -21 production for optimal help to B cells. These data illustrate complexities of SAP-dependent SLAM family receptor signaling, revealing a prominent role for SLAM receptor ligation in IL-4 production by GC CD4 T cells but not in T(FH) cell and GC T(FH) cell differentiation.
Conflict of interest statement
The authors have no financial conflicts of interest.
Figures

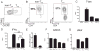
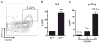




References
-
- Schoenberger SP, Crotty S. Immunologic memory. In: Paul WE, editor. Fundamental Immunology. 6. Lippincott Williams & Wilkins; Philadelphia: 2008. pp. 862–898.
-
- McHeyzer-Williams M. B Lymphocyte biology. In: Paul WE, editor. Fundamental Immunology. 6. Lippincott Williams & Wilkins; Philadelphia: 2008. pp. 289–311.
-
- Amanna IJ, Slifka MK, Crotty S. Immunity and immunological memory following smallpox vaccination. Immunol Rev. 2006;211:320–337. - PubMed
-
- Radbruch A, Muehlinghaus G, Luger EO, Inamine A, Smith KG, Dörner T, Hiepe F. Competence and competition: the challenge of becoming a long-lived plasma cell. Nat Rev Immunol. 2006;6:741–750. - PubMed
-
- Tangye SG, Tarlinton DM. Memory B cells: effectors of long-lived immune responses. Eur J Immunol. 2009;39:2065–2075. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

