FoxM1 mediates resistance to herceptin and paclitaxel
- PMID: 20530690
- PMCID: PMC2893542
- DOI: 10.1158/0008-5472.CAN-10-0545
FoxM1 mediates resistance to herceptin and paclitaxel
Abstract
Inherent and acquired therapeutic resistance in breast cancer remains a major clinical challenge. In human breast cancer samples, overexpression of the oncogenic transcription factor FoxM1 has been suggested to be a marker of poor prognosis. In this study, we report that FoxM1 overexpression confers resistance to the human epidermal growth factor receptor 2 monoclonal antibody Herceptin and microtubule-stabilizing drug paclitaxel, both as single agents and in combination. FoxM1 altered microtubule dynamics to protect tumor cells from paclitaxel-induced apoptosis. Mechanistic investigations revealed that the tubulin-destabilizing protein Stathmin, whose expression also confers resistance to paclitaxel, is a direct transcriptional target of FoxM1. Significantly, attenuating FoxM1 expression by small interfering RNA or an alternate reading frame (ARF)-derived peptide inhibitor increased therapeutic sensitivity. Our findings indicate that targeting FoxM1 could relieve therapeutic resistance in breast cancer.
Figures




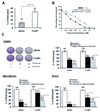

References
-
- Nakamura T, Furukawa Y, Nakagawa H, et al. Genome-wide cDNA microarray analysis of gene expression profiles in pancreatic cancers using populations of tumor cells and normal ductal epithelial cells selected for purity by laser microdissection. Oncogene. 2004;23:2385–2400. - PubMed
-
- Liu M, Dai B, Kang SH, et al. FoxM1B is overexpressed in human glioblastomas and critically regulates the tumorigenicity of glioma cells. Cancer Res. 2006;66:3593–3602. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

