Global coordination of transcriptional control and mRNA decay during cellular differentiation
- PMID: 20531409
- PMCID: PMC2913401
- DOI: 10.1038/msb.2010.38
Global coordination of transcriptional control and mRNA decay during cellular differentiation
Abstract
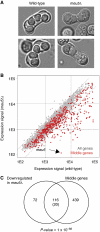
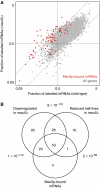
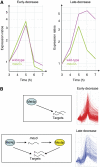
The function of transcription in dynamic gene expression programs has been extensively studied, but little is known about how it is integrated with RNA turnover at the genome-wide level. We investigated these questions using the meiotic gene expression program of Schizosaccharomyces pombe. We identified over 80 transcripts that co-purify with the meiotic-specific Meu5p RNA-binding protein. Their levels and half-lives were reduced in meu5 mutants, demonstrating that Meu5p stabilizes its targets. Most Meu5p-bound RNAs were also targets of the Mei4p transcription factor, which induces the transient expression of approximately 500 meiotic genes. Although many Mei4p targets showed sharp expression peaks, Meu5p targets had broad expression profiles. In the absence of meu5, all Mei4p targets were expressed with similar kinetics, indicating that Meu5p alters the global features of the gene expression program. As Mei4p activates meu5 transcription, Mei4p, Meu5p and their common targets form a feed-forward loop, a motif common in transcriptional networks but not studied in the context of mRNA decay. Our data provide insight into the topology of regulatory networks integrating transcriptional and posttranscriptional controls.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures


References
-
- Alon U (2007) Network motifs: theory and experimental approaches. Nat Rev Genet 8: 450–461 - PubMed
-
- Bähler J, Wu JQ, Longtine MS, Shah NG, McKenzie A III, Steever AB, Wach A, Philippsen P, Pringle JR (1998) Heterologous modules for efficient and versatile PCR-based gene targeting in Schizosaccharomyces pombe. Yeast 14: 943–951 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

