Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample
- PMID: 20534432
- PMCID: PMC3063599
- DOI: 10.1073/pnas.1000080107
Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample
Abstract
The ongoing revolution in high-throughput sequencing continues to democratize the ability of small groups of investigators to map the microbial component of the biosphere. In particular, the coevolution of new sequencing platforms and new software tools allows data acquisition and analysis on an unprecedented scale. Here we report the next stage in this coevolutionary arms race, using the Illumina GAIIx platform to sequence a diverse array of 25 environmental samples and three known "mock communities" at a depth averaging 3.1 million reads per sample. We demonstrate excellent consistency in taxonomic recovery and recapture diversity patterns that were previously reported on the basis of metaanalysis of many studies from the literature (notably, the saline/nonsaline split in environmental samples and the split between host-associated and free-living communities). We also demonstrate that 2,000 Illumina single-end reads are sufficient to recapture the same relationships among samples that we observe with the full dataset. The results thus open up the possibility of conducting large-scale studies analyzing thousands of samples simultaneously to survey microbial communities at an unprecedented spatial and temporal resolution.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



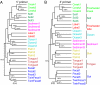

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

