Use of allostery to identify inhibitors of calmodulin-induced activation of Bacillus anthracis edema factor
- PMID: 20534570
- PMCID: PMC2895076
- DOI: 10.1073/pnas.0914611107
Use of allostery to identify inhibitors of calmodulin-induced activation of Bacillus anthracis edema factor
Abstract
Allostery plays a key role in the regulation of the activity and function of many biomolecules. And although many ligands act through allostery, no systematic use is made of it in drug design strategies. Here we describe a procedure for identifying the regions of a protein that can be used to control its activity through allostery. This procedure is based on the construction of a plausible conformational path, which describes protein transition between known active and inactive conformations. The path is calculated by using a framework approach that steers and markedly improves the conjugate peak refinement method. The evolution of conformations along this path was used to identify a putative allosteric site that could regulate activation of Bacillus anthracis adenylyl cyclase toxin (EF) by calmodulin. Conformations of the allosteric site at different steps along the path from the inactive (free) to the active (bound to calmodulin) forms of EF were used to perform virtual screenings and propose candidate EF inhibitors. Several candidates then proved to inhibit calmodulin-induced activation in an in vitro assay. The most potent compound fully inhibited EF at a concentration of 10 microM. The compounds also inhibited the related adenylyl cyclase toxin from Bordetella pertussis (CyaA). The specific homology between the putative allosteric sites in both toxins supports that these pockets are the actual binding sites of the selected inhibitors.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
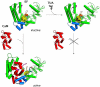
 in the inactive and active forms, respectively), EF and CaM are drawn in cartoon format. Switches A, B, and C in EF are highlighted in blue, orange, and magenta, respectively.
in the inactive and active forms, respectively), EF and CaM are drawn in cartoon format. Switches A, B, and C in EF are highlighted in blue, orange, and magenta, respectively.

References
-
- Kitchen D, Decornez H, Furr J, Bajorath J. Docking and scoring in virtual screening for drug discovery: Methods and applications. Nat Rev Drug Discov. 2004;3:935–949. - PubMed
-
- Pauli I, Timmers L, Caceres R, Soares M, de Azevedo W. In silico and in vitro: Identifying new drugs. Curr Drug Targets. 2008;9:1054–1061. - PubMed
-
- Rarey M, Kramer B, Lengauer T, Klebe G. A fast flexible docking method using an incremental construction algorithm. J Mol Biol. 1996;261:470–489. - PubMed
-
- Abagyan R, Totrov M, Kuznetsov D. ICM: A new method for protein modeling and design: Applications to docking and structure prediction from the distorted native conformation. J Comput Chem. 1994;15:488–506.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

