Three-amino acid spacing of presenilin endoproteolysis suggests a general stepwise cleavage of gamma-secretase-mediated intramembrane proteolysis
- PMID: 20534834
- PMCID: PMC6632680
- DOI: 10.1523/JNEUROSCI.1443-10.2010
Three-amino acid spacing of presenilin endoproteolysis suggests a general stepwise cleavage of gamma-secretase-mediated intramembrane proteolysis
Abstract
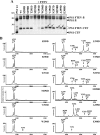
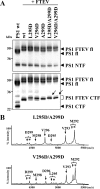
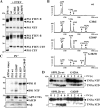
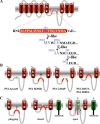
Presenilin (PS1 or PS2) is the catalytic component of the gamma-secretase complex, which mediates the final proteolytic processing step leading to the Alzheimer's disease (AD)-characterizing amyloid beta-peptide. PS is cleaved during complex assembly into its characteristic N- and C-terminal fragments. Both fragments are integral components of physiologically active gamma-secretase and harbor the two critical aspartyl residues of the active site domain. While the minimal subunit composition of gamma-secretase has been defined and numerous substrates were identified, the cellular mechanism of the endoproteolytic cleavage of PS is still unclear. We addressed this pivotal question by investigating whether familial AD (FAD)-associated PS1 mutations affect the precision of PS endoproteolysis in a manner similar to the way that such mutations shift the intramembrane cleavage of gamma-secretase substrates. We demonstrate that all FAD mutations investigated still allow endoproteolysis to occur. However, the precision of PS1 endoproteolysis is affected by PS1 mutations. Comparing the cleavage products generated by a variety of PS1 mutants revealed that specifically cleavages at positions 293 and 296 of PS1 are selectively affected. Systematic mutagenesis around the cleavage sites revealed a stepwise three amino acid spaced cleavage mechanism of PS endoproteolysis reminiscent to the epsilon-, zeta-, and gamma-cleavages described for typical gamma-secretase substrates, such as the beta-amyloid precursor protein. Our findings therefore suggest that intramembranous cleavage by gamma-secretase and related intramembrane-cleaving proteases may generally occur via stepwise endoproteolysis.
Figures



References
-
- Beher D, Wrigley JD, Nadin A, Evin G, Masters CL, Harrison T, Castro JL, Shearman MS. Pharmacological knock-down of the presenilin 1 heterodimer by a novel γ-secretase inhibitor: implications for presenilin biology. J Biol Chem. 2001;276:45394–45402. - PubMed
-
- Campbell WA, Iskandar MK, Reed ML, Xia W. Endoproteolysis of presenilin in vitro: inhibition by gamma-secretase inhibitors. Biochemistry. 2002;41:3372–3379. - PubMed
-
- Campbell WA, Reed ML, Strahle J, Wolfe MS, Xia W. Presenilin endoproteolysis mediated by an aspartyl protease activity pharmacologically distinct from gamma-secretase. J Neurochem. 2003;85:1563–1574. - PubMed
-
- Capell A, Saffrich R, Olivo JC, Meyn L, Walter J, Grünberg J, Mathews P, Nixon R, Dotti C, Haass C. Cellular expression and proteolytic processing of presenilin proteins is developmentally regulated during neuronal differentiation. J Neurochem. 1997;69:2432–2440. - PubMed
-
- Capell A, Grünberg J, Pesold B, Diehlmann A, Citron M, Nixon R, Beyreuther K, Selkoe DJ, Haass C. The proteolytic fragments of the Alzheimer's disease-associated presenilin-1 form heterodimers and occur as a 100–150-kDa molecular mass complex. J Biol Chem. 1998;273:3205–3211. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
