Comparative mitogenomics of Braconidae (Insecta: Hymenoptera) and the phylogenetic utility of mitochondrial genomes with special reference to Holometabolous insects
- PMID: 20537196
- PMCID: PMC2890569
- DOI: 10.1186/1471-2164-11-371
Comparative mitogenomics of Braconidae (Insecta: Hymenoptera) and the phylogenetic utility of mitochondrial genomes with special reference to Holometabolous insects
Abstract
Background: Animal mitochondrial genomes are potential models for molecular evolution and markers for phylogenetic and population studies. Previous research has shown interesting features in hymenopteran mitochondrial genomes. Here, we conducted a comparative study of mitochondrial genomes of the family Braconidae, one of the largest families of Hymenoptera, and assessed the utility of mitochondrial genomic data for phylogenetic inference at three different hierarchical levels, i.e., Braconidae, Hymenoptera, and Holometabola.
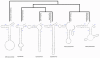
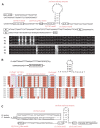
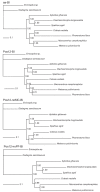
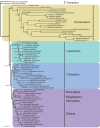
Results: Seven mitochondrial genomes from seven subfamilies of Braconidae were sequenced. Three of the four sequenced A+T-rich regions are shown to be inverted. Furthermore, all species showed reversal of strand asymmetry, suggesting that inversion of the A+T-rich region might be a synapomorphy of the Braconidae. Gene rearrangement events occurred in all braconid species, but gene rearrangement rates were not taxonomically correlated. Most rearranged genes were tRNAs, except those of Cotesia vestalis, in which 13 protein-coding genes and 14 tRNA genes changed positions or/and directions through three kinds of gene rearrangement events. Remote inversion is posited to be the result of two independent recombination events. Evolutionary rates were lower in species of the cyclostome group than those of noncyclostomes. Phylogenetic analyses based on complete mitochondrial genomes and secondary structure of rrnS supported a sister-group relationship between Aphidiinae and cyclostomes. Many well accepted relationships within Hymenoptera, such as paraphyly of Symphyta and Evaniomorpha, a sister-group relationship between Orussoidea and Apocrita, and monophyly of Proctotrupomorpha, Ichneumonoidea and Aculeata were robustly confirmed. New hypotheses, such as a sister-group relationship between Evanioidea and Aculeata, were generated. Among holometabolous insects, Hymenoptera was shown to be the sister to all other orders. Mecoptera was recovered as the sister-group of Diptera. Neuropterida (Neuroptera + Megaloptera), and a sister-group relationship with (Diptera + Mecoptera) were supported across all analyses.
Conclusions: Our comparative studies indicate that mitochondrial genomes are a useful phylogenetic tool at the ordinal level within Holometabola, at the superfamily within Hymenoptera and at the subfamily level within Braconidae. Variation at all of these hierarchical levels suggests that the utility of mitochondrial genomes is likely to be a valuable tool for systematics in other groups of arthropods.
Figures




References
-
- Zhang DX, Hewitt GM. Insect mitochondrial control region: a review of its structure, evolution and usefulness in evolutionary studies. Bio-chem Syst Ecol. 1997;25(2):99–120. doi: 10.1016/S0305-1978(96)00042-7. - DOI
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

