Combining isoelectric point-based fractionation, liquid chromatography and mass spectrometry to improve peptide detection and protein identification
- PMID: 20537905
- PMCID: PMC2927729
- DOI: 10.1016/j.jasms.2010.04.010
Combining isoelectric point-based fractionation, liquid chromatography and mass spectrometry to improve peptide detection and protein identification
Abstract
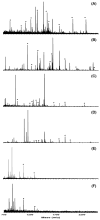
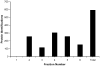
The off-line coupling of an isoelectric trapping device termed membrane separated wells for isoelectric focusing and trapping (MSWIFT) to mass spectrometry-based proteomic studies is described. The MSWIFT is a high capacity, high-throughput, mass spectrometry-compatible isoelectric trapping device that provides isoelectric point (pI)-based separations of complex mixtures of peptides. In MSWIFT, separation and analyte trapping are achieved by migrating the peptide ions through membranes having fixed pH values until the peptide pI is bracketed by the pH values of adjacent membranes. The pH values of the membranes can be tuned, thus affording a high degree of experimental flexibility. Specific advantages of using MSWIFT for sample prefractionation include: (1) small sample volumes (approximately 200 microL), (2) customized membranes over a large pH range, (3) flexibility in the number of desired fractions, (4) membrane compatibility with a variety of solvents systems, and (5) resulting fractions do not require sample cleanup before MS analysis. Here, we demonstrate the utility of MSWIFT for mass spectrometry-based detection of peptides in improving dynamic range and the reduction of ion suppression effects for high-throughput separations of tryptic peptides.
2010 American Society for Mass Spectrometry. Published by Elsevier Inc. All rights reserved.
Figures


References
-
- Righetti PG, Castagna A, Herbert B, Reymond F, Rossier JS. Prefractionation Techniques in Proteome Analysis. Proteomics. 2003;3:1397–1407. - PubMed
-
- Kolin A. Separation and Concentration of Proteins in a Ph Field Combined with an Electric Field. Journal of Chemical Physics. 1954;22:1628–1629.
-
- Svensson H. Isoelectric Fractionation, Analysis, and Characterization of Ampholytes in Natural Ph Gradients. 1. Differential Equation of Solute Concentrations at a Steady State and Its Solution for Simple Cases. Acta Chemica Scandinavica. 1961;15:325–341.
-
- Jensen PK, Pasa-Tolic L, Anderson GA, Horner JA, Lipton MS, Bruce JE, Smith RD. Probing Proteomes Using Capillary Isoelectric Focusing-Electrospray Ionization Fourier Transform Ion Cyclotron Resonance Mass Spectrometry. Analytical Chemistry. 1999;71:2076–2084. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

