Proteome-wide profiling of the MCF10AT breast cancer progression model
- PMID: 20543960
- PMCID: PMC2882958
- DOI: 10.1371/journal.pone.0011030
Proteome-wide profiling of the MCF10AT breast cancer progression model
Abstract
Background: Mapping the expression changes during breast cancer development should facilitate basic and translational research that will eventually improve our understanding and clinical management of cancer. However, most studies in this area are challenged by genetic and environmental heterogeneities associated with cancer.
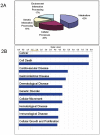
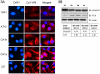
Methodology/principal findings: We conducted proteomics of the MCF10AT breast cancer model, which comprises of 4 isogenic xenograft-derived human cell lines that mimic different stages of breast cancer progression, using iTRAQ-based tandem mass spectrometry. Of more than 1200 proteins detected, 98 proteins representing at least 20 molecular function groups including kinases, proteases, adhesion, calcium binding and cytoskeletal proteins were found to display significant expression changes across the MCF10AT model. The number of proteins that showed different expression levels increased as disease progressed from AT1k pre-neoplastic cells to low grade CA1h cancer cells and high grade cancer cells. Bioinformatics revealed that MCF10AT model of breast cancer progression is associated with a major re-programming in metabolism, one of the first identified biochemical hallmarks of tumor cells (the "Warburg effect"). Aberrant expression of 3 novel breast cancer-associated proteins namely AK1, ATOX1 and HIST1H2BM were subsequently validated via immunoblotting of the MCF10AT model and immunohistochemistry of progressive clinical breast cancer lesions.
Conclusion/significance: The information generated by this study should serve as a useful reference for future basic and translational cancer research. Dysregulation of ATOX1, AK1 and HIST1HB2M could be detected as early as the pre-neoplastic stage. The findings have implications on early detection and stratification of patients for adjuvant therapy.
Conflict of interest statement
Figures


Comment in
-
Proteomic changes associated with breast cancer progression in the MCF10AT model.Pharmacogenomics. 2011 Jan;12(1):9-10. doi: 10.2217/pgs.10.194. Pharmacogenomics. 2011. PMID: 21174617 No abstract available.
Similar articles
-
Regulation of macrophage inhibitory factor (MIF) by epidermal growth factor receptor (EGFR) in the MCF10AT model of breast cancer progression.J Proteome Res. 2009 Aug;8(8):4062-76. doi: 10.1021/pr900430n. J Proteome Res. 2009. PMID: 19530702
-
Identification of differentially secreted biomarkers using LC-MS/MS in isogenic cell lines representing a progression of breast cancer.J Proteome Res. 2007 Aug;6(8):2993-3002. doi: 10.1021/pr060629m. Epub 2007 Jul 4. J Proteome Res. 2007. PMID: 17608509 Free PMC article.
-
Elevated NRD1 metalloprotease expression plays a role in breast cancer growth and proliferation.Genes Chromosomes Cancer. 2011 Oct;50(10):837-47. doi: 10.1002/gcc.20905. Epub 2011 Jul 18. Genes Chromosomes Cancer. 2011. PMID: 21769958
-
Delineating genetic alterations for tumor progression in the MCF10A series of breast cancer cell lines.PLoS One. 2010 Feb 15;5(2):e9201. doi: 10.1371/journal.pone.0009201. PLoS One. 2010. PMID: 20169162 Free PMC article.
-
Proteomic analysis of infiltrating ductal carcinoma tissues by coupled 2-D DIGE/MS/MS analysis.Mol Biol (Mosk). 2012 May-Jun;46(3):469-80. Mol Biol (Mosk). 2012. PMID: 22888636
Cited by
-
Multi-faceted quantitative proteomics analysis of histone H2B isoforms and their modifications.Epigenetics Chromatin. 2015 Apr 22;8:15. doi: 10.1186/s13072-015-0006-8. eCollection 2015. Epigenetics Chromatin. 2015. PMID: 25922622 Free PMC article.
-
Up-regulation of microRNA-145 associates with lymph node metastasis in colorectal cancer.PLoS One. 2014 Jul 14;9(7):e102017. doi: 10.1371/journal.pone.0102017. eCollection 2014. PLoS One. 2014. PMID: 25019299 Free PMC article.
-
Inhibition of Copper Transport Induces Apoptosis in Triple-Negative Breast Cancer Cells and Suppresses Tumor Angiogenesis.Mol Cancer Ther. 2019 May;18(5):873-885. doi: 10.1158/1535-7163.MCT-18-0667. Epub 2019 Mar 1. Mol Cancer Ther. 2019. PMID: 30824611 Free PMC article.
-
Analysis of the hippocampal proteome in ME7 prion disease reveals a predominant astrocytic signature and highlights the brain-restricted production of clusterin in chronic neurodegeneration.J Biol Chem. 2014 Feb 14;289(7):4532-45. doi: 10.1074/jbc.M113.502690. Epub 2013 Dec 23. J Biol Chem. 2014. PMID: 24366862 Free PMC article.
-
TrkA overexpression in non-tumorigenic human breast cell lines confers oncogenic and metastatic properties.Breast Cancer Res Treat. 2020 Feb;179(3):631-642. doi: 10.1007/s10549-019-05506-3. Epub 2019 Dec 10. Breast Cancer Res Treat. 2020. PMID: 31823098 Free PMC article.
References
-
- Santner SJ, Dawson PJ, Tait L, Soule HD, Eliason J, et al. Malignant MCF10CA1 cell lines derived from premalignant human breast epithelial MCF10AT cells. Breast Cancer Res Treat. 2001;65:101–110. - PubMed
-
- Strickland LB, Dawson PJ, Santner SJ, Miller FR. Progression of premalignant MCF10AT generates heterogeneous malignant variants with characteristic histologic types and immunohistochemical markers. Breast Cancer Res Treat. 2000;64:235–240. - PubMed
-
- Starcevic SL, Diotte NM, Zukowski KL, Cameron MJ, Novak RF. Oxidative DNA damage and repair in a cell lineage model of human proliferative breast disease (PBD). Toxicol Sci. 2003;75:74–81. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

