Shotgun proteomics identifies proteins specific for acute renal transplant rejection
- PMID: 20543976
- PMCID: PMC2883247
- DOI: 10.1002/prca.200900124
Shotgun proteomics identifies proteins specific for acute renal transplant rejection
Abstract
Purpose: Acute rejection (AR) remains the primary risk factor for renal transplant outcome; development of non-invasive diagnostic biomarkers for AR is an unmet need.
Experimental design: We used shotgun proteomics applying LC-MS/MS and ELISA to analyze a set of 92 urine samples, from patients with AR, stable grafts (STA), proteinuria (NS), and healthy controls.
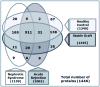
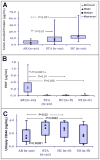
Results: A total of 1446 urinary proteins (UP) were identified along with a number of nonspecific proteinuria-specific, renal transplantation specific and AR-specific proteins. Relative abundance of identified UP was measured by protein-level spectral counts adopting a weighted fold-change statistic, assigning increased weight for more frequently observed proteins. We have identified alterations in a number of specific UP in AR, primarily relating to MHC antigens, the complement cascade and extra-cellular matrix proteins. A subset of proteins (uromodulin, SERPINF1 and CD44), have been further cross-validated by ELISA in an independent set of urine samples, for significant differences in the abundance of these UP in AR.
Conclusions and clinical relevance: This label-free, semi-quantitative approach for sampling the urinary proteome in normal and disease states provides a robust and sensitive method for detection of UP for serial, non-invasive clinical monitoring for graft rejection after kidney transplantation.
Keywords: Acute rejection; Biomarkers; ELISA; Renal transplantation; Urinary proteomics.
Copyright © 2010 WILEY-VCH Verlag GmbH & Co. KGaA, Weinheim.
Figures


References
-
- Cravatt BF, Simon GM, Yates JR., 3rd The biological impact of mass-spectrometry-based proteomics. Nature. 2007;450(7172):991–1000. - PubMed
-
- de Hoog CL, Mann M. Proteomics. Annu Rev Genomics Hum Genet. 2004;5:267–93. - PubMed
-
- Veenstra TD, et al. Biomarkers: mining the biofluid proteome. Mol Cell Proteomics. 2005;4(4):409–18. - PubMed
-
- Laronga C, Drake RR. Proteomic approach to breast cancer. Cancer Control. 2007;14(4):360–8. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

