Vibrio cholerae El Tor TcpA crystal structure and mechanism for pilus-mediated microcolony formation
- PMID: 20545841
- PMCID: PMC2939829
- DOI: 10.1111/j.1365-2958.2010.07244.x
Vibrio cholerae El Tor TcpA crystal structure and mechanism for pilus-mediated microcolony formation
Abstract
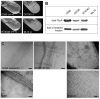
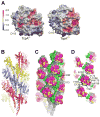
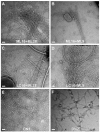
Type IV pili (T4P) are critical to virulence for Vibrio cholerae and other bacterial pathogens. Among their diverse functions, T4P mediate microcolony formation, which protects the bacteria from host defences and concentrates secreted toxins. The T4P of the two V. cholerae O1 disease biotypes, classical and El Tor, share 81% identity in their TcpA subunits, yet these filaments differ in their interaction patterns as assessed by electron microscopy. To understand the molecular basis for pilus-mediated microcolony formation, we solved a 1.5 A resolution crystal structure of N-terminally truncated El Tor TcpA and compared it with that of classical TcpA. Residues that differ between the two pilins are located on surface-exposed regions of the TcpA subunits. By iteratively changing these non-conserved amino acids in classical TcpA to their respective residues in El Tor TcpA, we identified residues that profoundly affect pilus:pilus interaction patterns and bacterial aggregation. These residues lie on either the protruding d-region of the TcpA subunit or in a cavity between pilin subunits in the pilus filament. Our results support a model whereby pili interact via intercalation of surface protrusions on one filament into depressions between subunits on adjacent filaments as a means to hold V. cholerae cells together in microcolonies.
Figures


References
-
- Barken KB, Pamp SJ, Yang L, Gjermansen M, Bertrand JJ, Klausen M, et al. Roles of type IV pili, flagellum-mediated motility and extracellular DNA in the formation of mature multicellular structures in Pseudomonas aeruginosa biofilms. Environ Microbiol. 2008;10:2331–2343. - PubMed
-
- Bieber D, Ramer SW, Wu CY, Murray WJ, Tobe T, Fernandez R, Schoolnik GK. Type IV pili, transient bacterial aggregates, and virulence of enteropathogenic Escherichia coli. Science. 1998;280:2114–2118. - PubMed
-
- Brunger AT, Adams PD, Clore GM, DeLano WL, Gros P, Grosse-Kunstleve RW, et al. Crystallography and NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr D Biol Crystallogr. 1998;54(Pt 5):905–921. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources

