Evaluation of a DNA microarray, the check-points ESBL/KPC array, for rapid detection of TEM, SHV, and CTX-M extended-spectrum beta-lactamases and KPC carbapenemases
- PMID: 20547813
- PMCID: PMC2916358
- DOI: 10.1128/AAC.01298-09
Evaluation of a DNA microarray, the check-points ESBL/KPC array, for rapid detection of TEM, SHV, and CTX-M extended-spectrum beta-lactamases and KPC carbapenemases
Abstract
Extended-spectrum beta-lactamases (ESBLs) and Klebsiella pneumoniae carbapenemases (KPC carbepenemases) have rapidly emerged worldwide and require rapid identification. The Check-Points ESBL/KPC array, a new commercial system based on genetic profiling for the direct identification of ESBL producers (SHV, TEM, and CTX-M) and of KPC producers, was evaluated. Well-characterized Gram-negative rods (Enterobacteriaceae, Pseudomonas aeruginosa, Acinetobacter baumannii) expressing various ss-lactamases (KPC-2, SHV, TEM, and CTX-M types) were used as well as wild-type reference strains and isolates harboring ss-lactamase genes not detected by the assay. In addition, phenotypically confirmed ESBL producers isolated in clinical samples over a 3-month period at the Bicetre hospital were analyzed using the Check-Points ESBL/KPC array and by standard PCR. The Check-Points ESBL/KPC array allowed fast detection of all TEM, SHV, and CTX-M ESBL genes and of the KPC-2 gene. The assay allowed easy differentiation between non-ESBL TEM and SHV and their ESBL derivatives. None of the other tested ss-lactamase genes were detected, underlining its high specificity. The technique is suited for Enterobacteriaceae but also for P. aeruginosa and A. baumannii. However, for nonfermenters, especially P. aeruginosa, a 1:10 dilution of the total DNA was necessary to detect KPC-2 and SHV-2a genes reliably. The Check-Points ESBL/KPC array is a powerful high-throughput tool for rapid identification of ESBLs and KPC producers in cultures. It provided definitive results within the same working day, allowing rapid implementation of isolation measures and appropriate antibiotic treatment. It showed an interesting potential for routine laboratory testing.
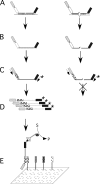
Figures

Similar articles
-
Evaluation of a DNA microarray (Check-MDR CT102) for rapid detection of TEM, SHV, and CTX-M extended-spectrum β-lactamases and of KPC, OXA-48, VIM, IMP, and NDM-1 carbapenemases.J Clin Microbiol. 2011 Apr;49(4):1608-13. doi: 10.1128/JCM.02607-10. Epub 2011 Feb 16. J Clin Microbiol. 2011. PMID: 21325547 Free PMC article.
-
Evaluation of a DNA microarray for the rapid detection of extended-spectrum β-lactamases (TEM, SHV and CTX-M), plasmid-mediated cephalosporinases (CMY-2-like, DHA, FOX, ACC-1, ACT/MIR and CMY-1-like/MOX) and carbapenemases (KPC, OXA-48, VIM, IMP and NDM).J Antimicrob Chemother. 2012 Aug;67(8):1865-9. doi: 10.1093/jac/dks156. Epub 2012 May 17. J Antimicrob Chemother. 2012. PMID: 22604450
-
Evaluation of a commercial microarray system for detection of SHV-, TEM-, CTX-M-, and KPC-type beta-lactamase genes in Gram-negative isolates.J Clin Microbiol. 2010 Jul;48(7):2618-22. doi: 10.1128/JCM.00568-10. Epub 2010 May 26. J Clin Microbiol. 2010. PMID: 20504993 Free PMC article.
-
The molecular basis of β-lactamase production in Gram-negative bacteria from Saudi Arabia.J Med Microbiol. 2015 Feb;64(Pt 2):127-136. doi: 10.1099/jmm.0.077834-0. Epub 2014 Nov 23. J Med Microbiol. 2015. PMID: 25418734 Review.
-
New definitions of extended-spectrum β-lactamase conferring worldwide emerging antibiotic resistance.Med Res Rev. 2012 Jan;32(1):216-32. doi: 10.1002/med.20210. Epub 2010 Jun 23. Med Res Rev. 2012. PMID: 20577973 Review.
Cited by
-
Evaluation of the NucliSENS EasyQ KPC assay for detection of Klebsiella pneumoniae carbapenemase-producing Enterobacteriaceae.J Clin Microbiol. 2013 Jun;51(6):1948-50. doi: 10.1128/JCM.00057-13. Epub 2013 Apr 3. J Clin Microbiol. 2013. PMID: 23554195 Free PMC article.
-
Evaluation of a DNA microarray (Check-MDR CT102) for rapid detection of TEM, SHV, and CTX-M extended-spectrum β-lactamases and of KPC, OXA-48, VIM, IMP, and NDM-1 carbapenemases.J Clin Microbiol. 2011 Apr;49(4):1608-13. doi: 10.1128/JCM.02607-10. Epub 2011 Feb 16. J Clin Microbiol. 2011. PMID: 21325547 Free PMC article.
-
Evaluation of a New Commercial Microarray Platform for the Simultaneous Detection of β-Lactamase and mcr-1 and mcr-2 Genes in Enterobacteriaceae.J Clin Microbiol. 2017 Oct;55(10):3138-3141. doi: 10.1128/JCM.01056-17. Epub 2017 Aug 2. J Clin Microbiol. 2017. PMID: 28768732 Free PMC article. No abstract available.
-
Mechanisms of Resistance in Gram-Negative Urinary Pathogens: From Country-Specific Molecular Insights to Global Clinical Relevance.Diagnostics (Basel). 2021 Apr 28;11(5):800. doi: 10.3390/diagnostics11050800. Diagnostics (Basel). 2021. PMID: 33925181 Free PMC article. Review.
-
Emerging rapid resistance testing methods for clinical microbiology laboratories and their potential impact on patient management.Biomed Res Int. 2014;2014:375681. doi: 10.1155/2014/375681. Epub 2014 Sep 17. Biomed Res Int. 2014. PMID: 25343142 Free PMC article. Review.
References
-
- Arlet, G., G. Brami, D. Decre, A. Flippo, O. Gaillot, P. H. Lagrange, and A. Phillipon. 1995. Molecular characterization by PCR-restriction fragment length polymorphism of TEM ß-lactamases. FEMS Microbiol. Lett. 134:203-208. - PubMed
-
- Bedenic, B., J. Vranes, L. J. Mihaljevic, M. Tonkic, M. Sviben, V. Plecko, and S. Kalenic. 2007. Sensitivity and specificity of various ß-lactam antibiotics and phenotypical methods for detection of TEM, SHV and CTX-M extended-spectrum ß-lactamases. J. Chemother. 19:127-139. - PubMed
-
- Birkett, C. I., H. A. Ludlam, N. Woodford, D. F. Brown, N. M. Brown, M. T. Roberts, N. Milner, and M. D. Curran. 2007. Real-time TaqMan PCR for rapid detection and typing of genes encoding CTX-M extended-spectrum ß-lactamases. J. Med. Microbiol. 56:52-55. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

