Globular state in the oligomers formed by Abeta peptides
- PMID: 20550420
- PMCID: PMC2896418
- DOI: 10.1063/1.3447894
Globular state in the oligomers formed by Abeta peptides
Abstract
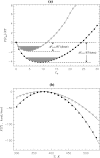
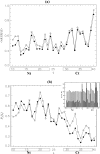
Replica exchange molecular dynamics and implicit solvent model are used to study two oligomeric species of Abeta peptides, dimer and tetramer, which are typically observed in in vitro experiments. Based on the analysis of free energy landscapes, density distributions, and chain flexibility, we propose that the oligomer formation is a continuous transition occurring without metastable states. The density distribution computations suggest that Abeta oligomer consists of two volume regions-the core with fairly flat density profile and the surface layer with rapidly decreasing density. The core is mostly formed by the N-terminal residues, whereas the C-terminal tends to occur in the surface layer. Lowering the temperature results in the redistribution of peptide atoms from the surface layer into the core. Using these findings, we argue that Abeta oligomer resembles polymer globule in poor solvent. Abeta dimers and tetramers are found to be structurally similar suggesting that the conformations of Abeta peptides do not depend on the order of small oligomers.
Figures



Similar articles
-
Mapping conformational ensembles of aβ oligomers in molecular dynamics simulations.Biophys J. 2010 Sep 22;99(6):1949-58. doi: 10.1016/j.bpj.2010.07.008. Biophys J. 2010. PMID: 20858441 Free PMC article.
-
Oligomer Formation of Amyloid-β(29-42) from Its Monomers Using the Hamiltonian Replica-Permutation Molecular Dynamics Simulation.J Phys Chem B. 2016 Jul 14;120(27):6555-61. doi: 10.1021/acs.jpcb.6b03828. Epub 2016 Jun 24. J Phys Chem B. 2016. PMID: 27281682
-
Conformational ensemble and polymorphism of the all-atom Alzheimer's Aβ(37-42) amyloid peptide oligomers.J Phys Chem B. 2013 May 16;117(19):5831-40. doi: 10.1021/jp401563n. Epub 2013 May 6. J Phys Chem B. 2013. PMID: 23581814
-
Understanding amyloid fibril nucleation and aβ oligomer/drug interactions from computer simulations.Acc Chem Res. 2014 Feb 18;47(2):603-11. doi: 10.1021/ar4002075. Epub 2013 Dec 24. Acc Chem Res. 2014. PMID: 24368046 Review.
-
Evolutionary, physicochemical, and functional mechanisms of protein homooligomerization.Prog Mol Biol Transl Sci. 2013;117:3-24. doi: 10.1016/B978-0-12-386931-9.00001-5. Prog Mol Biol Transl Sci. 2013. PMID: 23663963 Free PMC article. Review.
Cited by
-
Survey of the Aβ-peptide structural diversity: molecular dynamics approaches.Biophys Rev. 2024 Nov 20;16(6):701-722. doi: 10.1007/s12551-024-01253-y. eCollection 2024 Dec. Biophys Rev. 2024. PMID: 39830132 Review.
-
Mapping conformational ensembles of aβ oligomers in molecular dynamics simulations.Biophys J. 2010 Sep 22;99(6):1949-58. doi: 10.1016/j.bpj.2010.07.008. Biophys J. 2010. PMID: 20858441 Free PMC article.
-
Polymorphic structures of Alzheimer's β-amyloid globulomers.PLoS One. 2011;6(6):e20575. doi: 10.1371/journal.pone.0020575. Epub 2011 Jun 7. PLoS One. 2011. PMID: 21687730 Free PMC article.
-
Naproxen interferes with the assembly of Aβ oligomers implicated in Alzheimer's disease.Biophys J. 2011 Apr 20;100(8):2024-32. doi: 10.1016/j.bpj.2011.02.044. Biophys J. 2011. PMID: 21504739 Free PMC article.
-
Molecular interactions of Alzheimer's biomarker FDDNP with Aβ peptide.Biophys J. 2012 Dec 5;103(11):2341-51. doi: 10.1016/j.bpj.2012.10.003. Biophys J. 2012. PMID: 23283233 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

