Helix unwinding and base flipping enable human MTERF1 to terminate mitochondrial transcription
- PMID: 20550934
- PMCID: PMC2887341
- DOI: 10.1016/j.cell.2010.05.018
Helix unwinding and base flipping enable human MTERF1 to terminate mitochondrial transcription
Abstract
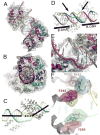
Defects in mitochondrial gene expression are associated with aging and disease. Mterf proteins have been implicated in modulating transcription, replication and protein synthesis. We have solved the structure of a member of this family, the human mitochondrial transcriptional terminator MTERF1, bound to dsDNA containing the termination sequence. The structure indicates that upon sequence recognition MTERF1 unwinds the DNA molecule, promoting eversion of three nucleotides. Base flipping is critical for stable binding and transcriptional termination. Additional structural and biochemical results provide insight into the DNA binding mechanism and explain how MTERF1 recognizes its target sequence. Finally, we have demonstrated that the mitochondrial pathogenic G3249A and G3244A mutations interfere with key interactions for sequence recognition, eliminating termination. Our results provide insight into the role of mterf proteins and suggest a link between mitochondrial disease and the regulation of mitochondrial transcription.
Copyright 2010 Elsevier Inc. All rights reserved.
Figures
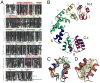
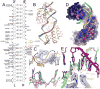



Comment in
-
MTERF1 gives mtDNA an unusual twist.Cell Metab. 2010 Jul 7;12(1):3-4. doi: 10.1016/j.cmet.2010.06.006. Cell Metab. 2010. PMID: 20620988
Similar articles
-
Mitochondrial transcription: how does it end?Transcription. 2011 Jan-Feb;2(1):32-6. doi: 10.4161/trns.2.1.14006. Transcription. 2011. PMID: 21326908 Free PMC article.
-
Mitochondrial transcription termination factor 1 directs polar replication fork pausing.Nucleic Acids Res. 2016 Jul 8;44(12):5732-42. doi: 10.1093/nar/gkw302. Epub 2016 Apr 25. Nucleic Acids Res. 2016. PMID: 27112570 Free PMC article.
-
Base Flipping by MTERF1 Can Accommodate Multiple Conformations and Occurs in a Stepwise Fashion.J Mol Biol. 2016 Jun 19;428(12):2542-2556. doi: 10.1016/j.jmb.2015.10.021. Epub 2015 Oct 30. J Mol Biol. 2016. PMID: 26523681 Free PMC article.
-
The MTERF family proteins: mitochondrial transcription regulators and beyond.Biochim Biophys Acta. 2009 May;1787(5):303-11. doi: 10.1016/j.bbabio.2009.01.013. Epub 2009 Jan 30. Biochim Biophys Acta. 2009. PMID: 19366610 Review.
-
Hitting the brakes: termination of mitochondrial transcription.Biochim Biophys Acta. 2012 Sep-Oct;1819(9-10):939-47. doi: 10.1016/j.bbagrm.2011.11.004. Epub 2011 Nov 25. Biochim Biophys Acta. 2012. PMID: 22137970 Free PMC article. Review.
Cited by
-
Arabidopsis thaliana mTERF proteins: evolution and functional classification.Front Plant Sci. 2012 Oct 15;3:233. doi: 10.3389/fpls.2012.00233. eCollection 2012. Front Plant Sci. 2012. PMID: 23087700 Free PMC article.
-
Structure, mechanism, and regulation of mitochondrial DNA transcription initiation.J Biol Chem. 2020 Dec 25;295(52):18406-18425. doi: 10.1074/jbc.REV120.011202. Epub 2020 Oct 30. J Biol Chem. 2020. PMID: 33127643 Free PMC article. Review.
-
Pathological and Therapeutic Advances in Parkinson's Disease: Mitochondria in the Interplay.J Alzheimers Dis. 2023;94(s1):S399-S428. doi: 10.3233/JAD-220682. J Alzheimers Dis. 2023. PMID: 36093711 Free PMC article. Review.
-
Mitochondrial Gene Expression and Beyond-Novel Aspects of Cellular Physiology.Cells. 2019 Dec 19;9(1):17. doi: 10.3390/cells9010017. Cells. 2019. PMID: 31861673 Free PMC article. Review.
-
Energy Landscapes for Base-Flipping in a Model DNA Duplex.J Phys Chem B. 2022 Apr 28;126(16):3012-3028. doi: 10.1021/acs.jpcb.2c00340. Epub 2022 Apr 15. J Phys Chem B. 2022. PMID: 35427136 Free PMC article.
References
-
- Anderson S, Bankier AT, Barrell BG, et al. Sequence and organization of the human mitochondrial genome. Nature. 1981;290:457–465. - PubMed
-
- Asin-Cayuela J, Schwend T, Farge G, Gustafsson CM. The human mitochondrial transcription termination factor (mTERF) is fully active in vitro in the non-phosphorylated form. J Biol Chem. 2005;280:25499–25505. - PubMed
-
- Asin-Cayuela J, Gustafsson CM. Mitochondrial transcription and its regulation in mammalian cells. Trends Biochem Sci. 2007;32:111–117. - PubMed
-
- Cowtan K. Joint CCP4 and ESF-EACMB Newsletter on Protein. Crystallography. 1994;31:34–38.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

