Novel phorbol ester-binding motif mediates hormonal activation of Na+/H+ exchanger
- PMID: 20551318
- PMCID: PMC2924106
- DOI: 10.1074/jbc.M110.130120
Novel phorbol ester-binding motif mediates hormonal activation of Na+/H+ exchanger
Abstract
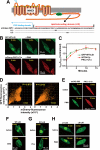
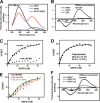
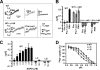
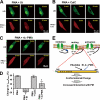
Protein kinase C (PKC) is considered crucial for hormonal Na(+)/H(+) exchanger (NHE1) activation because phorbol esters (PEs) strongly activate NHE1. However, here we report that rather than PKC, direct binding of PEs/diacylglycerol to the NHE1 lipid-interacting domain (LID) and the subsequent tighter association of LID with the plasma membrane mainly underlies NHE1 activation. We show that (i) PEs directly interact with the LID of NHE1 in vitro, (ii) like PKC, green fluorescent protein (GFP)-labeled LID translocates to the plasma membrane in response to PEs and receptor agonists, (iii) LID mutations markedly inhibit these interactions and PE/receptor agonist-induced NHE1 activation, and (iv) PKC inhibitors ineffectively block NHE1 activation, except staurosporin, which itself inhibits NHE1 via LID. Thus, we propose a PKC-independent mechanism of NHE1 regulation via a PE-binding motif previously unrecognized.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

