Inferring bacterial genome flux while considering truncated genes
- PMID: 20551435
- PMCID: PMC2940306
- DOI: 10.1534/genetics.110.118448
Inferring bacterial genome flux while considering truncated genes
Abstract
Bacterial gene content variation during the course of evolution has been widely acknowledged and its pattern has been actively modeled in recent years. Gene truncation or gene pseudogenization also plays an important role in shaping bacterial genome content. Truncated genes could also arise from small-scale lateral gene transfer events. Unfortunately, the information of truncated genes has not been considered in any existing mathematical models on gene content variation. In this study, we developed a model to incorporate truncated genes. Maximum-likelihood estimates (MLEs) of the new model reveal fast rates of gene insertions/deletions on recent branches, suggesting a fast turnover of many recently transferred genes. The estimates also suggest that many truncated genes are in the process of being eliminated from the genome. Furthermore, we demonstrate that the ignorance of truncated genes in the estimation does not lead to a systematic bias but rather has a more complicated effect. Analysis using the new model not only provides more accurate estimates on gene gains/losses (or insertions/deletions), but also reduces any concern of a systematic bias from applying simplified models to bacterial genome evolution. Although not a primary purpose, the model incorporating truncated genes could be potentially used for phylogeny reconstruction using gene family content.
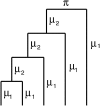
Figures


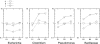

Similar articles
-
High rates of lateral gene transfer are not due to false diagnosis of gene absence.Gene. 2008 Sep 15;421(1-2):27-31. doi: 10.1016/j.gene.2008.06.015. Epub 2008 Jun 14. Gene. 2008. PMID: 18601986
-
Uncovering rate variation of lateral gene transfer during bacterial genome evolution.BMC Genomics. 2008 May 20;9:235. doi: 10.1186/1471-2164-9-235. BMC Genomics. 2008. PMID: 18492275 Free PMC article.
-
Does gene translocation accelerate the evolution of laterally transferred genes?Genetics. 2009 Aug;182(4):1365-75. doi: 10.1534/genetics.109.104216. Epub 2009 May 27. Genetics. 2009. PMID: 19474197 Free PMC article.
-
Deletional bias and the evolution of bacterial genomes.Trends Genet. 2001 Oct;17(10):589-96. doi: 10.1016/s0168-9525(01)02447-7. Trends Genet. 2001. PMID: 11585665 Review.
-
Models, algorithms and programs for phylogeny reconciliation.Brief Bioinform. 2011 Sep;12(5):392-400. doi: 10.1093/bib/bbr045. Brief Bioinform. 2011. PMID: 21949266 Review.
Cited by
-
Inferring horizontal gene transfer.PLoS Comput Biol. 2015 May 28;11(5):e1004095. doi: 10.1371/journal.pcbi.1004095. eCollection 2015 May. PLoS Comput Biol. 2015. PMID: 26020646 Free PMC article.
-
Complete Genome Analysis of Thermus parvatiensis and Comparative Genomics of Thermus spp. Provide Insights into Genetic Variability and Evolution of Natural Competence as Strategic Survival Attributes.Front Microbiol. 2017 Jul 27;8:1410. doi: 10.3389/fmicb.2017.01410. eCollection 2017. Front Microbiol. 2017. PMID: 28798737 Free PMC article.
-
Comparative Genome Analysis of 19 Trueperella pyogenes Strains Originating from Different Animal Species Reveal a Genetically Diverse Open Pan-Genome.Antibiotics (Basel). 2022 Dec 24;12(1):24. doi: 10.3390/antibiotics12010024. Antibiotics (Basel). 2022. PMID: 36671226 Free PMC article.
-
Homologous recombination drives both sequence diversity and gene content variation in Neisseria meningitidis.Genome Biol Evol. 2013;5(9):1611-27. doi: 10.1093/gbe/evt116. Genome Biol Evol. 2013. PMID: 23902748 Free PMC article.
-
Horizontal transfer and gene conversion as an important driving force in shaping the landscape of mitochondrial introns.G3 (Bethesda). 2014 Apr 16;4(4):605-12. doi: 10.1534/g3.113.009910. G3 (Bethesda). 2014. PMID: 24515269 Free PMC article.
References
-
- Archibald, J. M., and A. J. Roger, 2002. Gene duplication and gene conversion shape the evolution of archaeal chaperonins. J. Mol. Biol. 316 1041–1050. - PubMed
-
- Berg, O. G., and C. G. Kurland, 2002. Evolution of microbial genomes: sequence acquisition and loss. Mol. Biol. Evol. 19 2265–2276. - PubMed
-
- Brochier, C., H. Philippe and D. Moreira, 2000. The evolutionary history of ribosomal protein RpS14: horizontal gene transfer at the heart of the ribosome. Trends Genet. 16 529–533. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

