The structure of the TNFRSF13C promoter enables differential expression of BAFF-R during B cell ontogeny and terminal differentiation
- PMID: 20554963
- PMCID: PMC3780588
- DOI: 10.4049/jimmunol.1001120
The structure of the TNFRSF13C promoter enables differential expression of BAFF-R during B cell ontogeny and terminal differentiation
Erratum in
- J Immunol. 2010 Nov 15;185(10):6384
Abstract
The B cell-activating factor of the TNF family receptor (BAFF-R), encoded by the TNFRSF13C gene, is critically important for transitional B cell survival to maturity. Thus, ligation of BAFF-R by BAFF delivers a potent survival signal. Reports implicating the BAFF/BAFF-R signaling axis in the pathogenesis of autoimmune human diseases and B lineage malignancies have largely prompted studies focusing on BAFF expression; however, there is an equally critical need to better understand BAFF-R expression. Initial BAFF-R expression, although characterized in murine B cells, has not yet been reported in human B lymphopoiesis. In this study, we first demonstrate that BAFF-R expression is absent from early precursors and is acquired by bone marrow B cells newly expressing the BCR. We next focused on identifying the specific genomic region that controls BAFF-R expression in mature B cells (i.e., the TNFRSF13C promoter). To accomplish this, we used in silico tools examining interspecies genomic conservation in conjunction with reporter constructs transfected into malignant B and plasma cell lines. DNase protection assays using nuclear extracts from BAFF-R-expressing cells suggested potential regulatory sites, which allowed the generation of EMSA probes that bound NFs specific to BAFF-R-expressing cells. With a more stringent analysis of interspecies homology, these assays identified a site at which a single nucleotide substitution could distinctly impact promoter activity. Finally, chromatin immunoprecipitation assays revealed the in vivo binding of the specific transcription factor c-Rel to the most proximal genomic region, and c-Rel small interfering RNA transfections in BAFF-R-expressing lines demonstrated a coincident knockdown of both c-Rel and BAFF-R mRNA.
Conflict of interest statement
The authors have no financial conflict of interest to disclose.
Figures
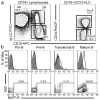






References
-
- Moore PA, Belvedere O, Orr A, Pieri K, LaFleur DW, Feng P, Soppet D, Charters M, Gentz R, Parmelee D, Li Y, Galperina O, Giri J, Roschke V, Nardelli B, Carrell J, Sosnovtseva S, Greenfield W, Ruben SM, Olsen HS, Fikes J, Hilbert DM. BLyS: member of the tumor necrosis factor family and B lymphocyte stimulator. Science. 1999;285:260–263. - PubMed
-
- Schneider P, MacKay F, Steiner V, Hofmann K, Bodmer JL, Holler N, Ambrose C, Lawton P, Bixler S, Acha-Orbea H, Valmori D, Romero P, Werner-Favre C, Zubler RH, Browning JL, Tschopp J. BAFF, a novel ligand of the tumor necrosis factor family, stimulates B cell growth. J Exp Med. 1999;189:1747–1756. - PMC - PubMed
-
- Yan M, Brady JR, Chan B, Lee WP, Hsu B, Harless S, Cancro M, Grewal IS, Dixit VM. Identification of a novel receptor for B lymphocyte stimulator that is mutated in a mouse strain with severe B cell deficiency. Curr Biol. 2001;11:1547–1552. - PubMed
-
- Schiemann B, Gommerman JL, Vora K, Cachero TG, Shulga-Morskaya S, Dobles M, Frew E, Scott ML. An essential role for BAFF in the normal development of B cells through a BCMA-independent pathway.[see comment] Science. 2001;293:2111–2114. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

