Thermodynamics of RNA duplexes modified with unlocked nucleic acid nucleotides
- PMID: 20562222
- PMCID: PMC2965255
- DOI: 10.1093/nar/gkq561
Thermodynamics of RNA duplexes modified with unlocked nucleic acid nucleotides
Abstract
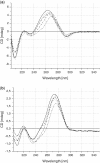
Thermodynamics provides insights into the influence of modified nucleotide residues on stability of nucleic acids and is crucial for designing duplexes with given properties. In this article, we introduce detailed thermodynamic analysis of RNA duplexes modified with unlocked nucleic acid (UNA) nucleotide residues. We investigate UNA single substitutions as well as model mismatch and dangling end effects. UNA residues placed in a central position makes RNA duplex structure less favourable by 4.0-6.6 kcal/mol. Slight destabilization, by ∼0.5-1.5 kcal/mol, is observed for 5'- or 3'-terminal UNA residues. Furthermore, thermodynamic effects caused by UNA residues are extremely additive with ΔG°(37) conformity up to 98%. Direct mismatches involving UNA residues decrease the thermodynamic stability less than unmodified mismatches in RNA duplexes. Additionally, the presence of UNA residues adjacent to unpaired RNA residues reduces mismatch discrimination. Thermodynamic analysis of UNA 5'- and 3'-dangling ends revealed that stacking interactions of UNA residues are always less favourable than that of RNA residues. Finally, circular dichroism spectra imply no changes in overall A-form structure of UNA-RNA/RNA duplexes relative to the unmodified RNA duplexes.
Figures
Similar articles
-
The influence of various modified nucleotides placed as 3'-dangling end on thermal stability of RNA duplexes.Biophys Chem. 2002 Jun 19;97(2-3):243-9. doi: 10.1016/s0301-4622(02)00073-x. Biophys Chem. 2002. PMID: 12050013
-
The influence of locked nucleic acid residues on the thermodynamic properties of 2'-O-methyl RNA/RNA heteroduplexes.Nucleic Acids Res. 2005 Sep 9;33(16):5082-93. doi: 10.1093/nar/gki789. Print 2005. Nucleic Acids Res. 2005. PMID: 16155181 Free PMC article.
-
Long RNA dangling end has large energetic contribution to duplex stability.J Am Chem Soc. 2002 Sep 4;124(35):10367-72. doi: 10.1021/ja0255406. J Am Chem Soc. 2002. PMID: 12197739
-
Contribution of 3'T and 3'TT overhangs to the thermodynamic stability of model siRNA duplexes.Biophys Chem. 2019 Mar;246:35-39. doi: 10.1016/j.bpc.2018.12.006. Epub 2019 Jan 7. Biophys Chem. 2019. PMID: 30660935 Free PMC article.
-
Thermodynamic characterization of single mismatches found in naturally occurring RNA.Biochemistry. 2007 Nov 20;46(46):13425-36. doi: 10.1021/bi701311c. Epub 2007 Oct 24. Biochemistry. 2007. PMID: 17958380
Cited by
-
5' Unlocked Nucleic Acid Modification Improves siRNA Targeting.Mol Ther Nucleic Acids. 2013 Jul 2;2(7):e103. doi: 10.1038/mtna.2013.36. Mol Ther Nucleic Acids. 2013. PMID: 23820891 Free PMC article.
-
A systematic study on the influence of thermodynamic asymmetry of 5'-ends of siRNA duplexes in relation to their silencing potency.Sci Rep. 2019 Feb 21;9(1):2477. doi: 10.1038/s41598-018-36620-9. Sci Rep. 2019. PMID: 30792489 Free PMC article.
-
Watson-Crick-like pairs in CCUG repeats: evidence for tautomeric shifts or protonation.RNA. 2016 Jan;22(1):22-31. doi: 10.1261/rna.052399.115. Epub 2015 Nov 5. RNA. 2016. PMID: 26543073 Free PMC article.
-
Dissecting the chemical interactions and substrate structural signatures governing RNA polymerase II trigger loop closure by synthetic nucleic acid analogues.Nucleic Acids Res. 2014 May;42(9):5863-70. doi: 10.1093/nar/gku238. Epub 2014 Apr 1. Nucleic Acids Res. 2014. PMID: 24692664 Free PMC article.
-
Improved thrombin binding aptamer by incorporation of a single unlocked nucleic acid monomer.Nucleic Acids Res. 2011 Feb;39(3):1155-64. doi: 10.1093/nar/gkq823. Epub 2010 Sep 24. Nucleic Acids Res. 2011. PMID: 20870750 Free PMC article.
References
-
- Micklefield J. Backbone modification of nucleic acids: synthesis, structure and therapeutic applications. Curr. Med. Chem. 2001;8:1157–1179. - PubMed
-
- Doktycz MJ. Encyclopedia of Life Science. Chichester: John Wiley & Sons; 1997. Nucleic acids: thermal stability and denaturation; pp. 3123–3140.
-
- Turner DH, Sugimoto N, Kierzek R, Dreiker SD. Free energy increments for hydrogen-bonds in nucleic acid base pairs. J. Am. Chem. Soc. 1987;109:3783–3785.
-
- Kool ET. Hydrogen bonding, base stacking, and steric effects in DNA replication. Annu. Rev. Biophys. Biomol. Stuct. 2001;30:1–22. - PubMed

