Xer1-mediated site-specific DNA inversions and excisions in Mycoplasma agalactiae
- PMID: 20562305
- PMCID: PMC2937384
- DOI: 10.1128/JB.01537-09
Xer1-mediated site-specific DNA inversions and excisions in Mycoplasma agalactiae
Abstract
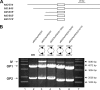
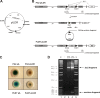
Surface antigen variation in Mycoplasma agalactiae, the etiologic agent of contagious agalactia in sheep and goats, is governed by site-specific recombination within the vpma multigene locus encoding the Vpma family of variable surface lipoproteins. This high-frequency Vpma phase switching was previously shown to be mediated by a Xer1 recombinase encoded adjacent to the vpma locus. In this study, it was demonstrated in Escherichia coli that the Xer1 recombinase is responsible for catalyzing vpma gene inversions between recombination sites (RS) located in the 5'-untranslated region (UTR) in all six vpma genes, causing cleavage and strand exchange within a 21-bp conserved region that serves as a recognition sequence. It was further shown that the outcome of the site-specific recombination event depends on the orientation of the two vpma RS, as direct or inverted repeats. While recombination between inverted vpma RS led to inversions, recombination between direct repeat vpma RS led to excisions. Using a newly developed excision assay based on the lacZ reporter system, we were able to successfully demonstrate under native conditions that such Xer1-mediated excisions can indeed also occur in the M. agalactiae type strain PG2, whereas they were not observed in the control xer1-disrupted VpmaY phase-locked mutant (PLMY), which lacks Xer1 recombinase. Unless there are specific regulatory mechanisms preventing such excisions, this might be the cost that the pathogen has to render at the population level for maintaining this high-frequency phase variation machinery.
Figures





Similar articles
-
Xer1-independent mechanisms of Vpma phase variation in Mycoplasma agalactiae are triggered by Vpma-specific antibodies.Int J Med Microbiol. 2017 Dec;307(8):443-451. doi: 10.1016/j.ijmm.2017.10.005. Epub 2017 Nov 6. Int J Med Microbiol. 2017. PMID: 29122515
-
Surface diversity in Mycoplasma agalactiae is driven by site-specific DNA inversions within the vpma multigene locus.J Bacteriol. 2002 Nov;184(21):5987-98. doi: 10.1128/JB.184.21.5987-5998.2002. J Bacteriol. 2002. PMID: 12374833 Free PMC article.
-
Phase-locked mutants of Mycoplasma agalactiae: defining the molecular switch of high-frequency Vpma antigenic variation.Mol Microbiol. 2008 Mar;67(6):1196-210. doi: 10.1111/j.1365-2958.2007.06103.x. Epub 2008 Jan 30. Mol Microbiol. 2008. PMID: 18248580 Free PMC article.
-
[Molecular basis of Mycoplasma agalactiae pathogenicity].Berl Munch Tierarztl Wochenschr. 2004 Nov-Dec;117(11-12):472-9. Berl Munch Tierarztl Wochenschr. 2004. PMID: 15584429 Review. German.
-
Structure and function of the shufflon in plasmid R64.Adv Biophys. 2004;38:183-213. Adv Biophys. 2004. PMID: 15493334 Review.
Cited by
-
Role of Vpma phase variation in Mycoplasma agalactiae pathogenesis.FEMS Immunol Med Microbiol. 2012 Dec;66(3):307-22. doi: 10.1111/j.1574-695X.2012.01010.x. Epub 2012 Aug 21. FEMS Immunol Med Microbiol. 2012. PMID: 22809092 Free PMC article.
-
Vpma phase variation is important for survival and persistence of Mycoplasma agalactiae in the immunocompetent host.PLoS Pathog. 2017 Sep 28;13(9):e1006656. doi: 10.1371/journal.ppat.1006656. eCollection 2017 Sep. PLoS Pathog. 2017. PMID: 28957426 Free PMC article.
-
Interaction of the putative tyrosine recombinases RipX (UU145), XerC (UU222), and CodV (UU529) of Ureaplasma parvum serovar 3 with specific DNA.FEMS Microbiol Lett. 2013 Mar;340(1):55-64. doi: 10.1111/1574-6968.12077. Epub 2013 Jan 31. FEMS Microbiol Lett. 2013. PMID: 23305333 Free PMC article.
-
Disruption of the pdhB pyruvate dehydrogenase [corrected] gene affects colony morphology, in vitro growth and cell invasiveness of Mycoplasma agalactiae.PLoS One. 2015 Mar 23;10(3):e0119706. doi: 10.1371/journal.pone.0119706. eCollection 2015. PLoS One. 2015. PMID: 25799063 Free PMC article.
-
Mycoplasma agalactiae ST35: a new sequence type with a minimal accessory genome primarily affecting goats.BMC Vet Res. 2022 Jan 11;18(1):29. doi: 10.1186/s12917-021-03128-w. BMC Vet Res. 2022. PMID: 35016679 Free PMC article.
References
-
- Barre, F. X., and D. J. Sherratt. 2002. Xer site-specific recombination: promoting chromosome segregation, p. 149-161. In N. L. Craig, R. Craigie, M. Gellert, and A. M. Lambowitz (ed.), Mobile DNA II. ASM Press, Washington, DC.
-
- Bayliss, C. D. 2009. Determinants of phase variation rate and the fitness implications of differing rates for bacterial pathogens and commensals. FEMS Microbiol. Rev. 33:504-520. - PubMed
-
- Bhugra, B., L. L. Voelker, N. Zou, H. Yu, and K. Dybvig. 1995. Mechanism of antigenic variation in Mycoplasma pulmonis: interwoven, site-specific DNA inversions. Mol. Microbiol. 18:703-714. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

