Epigenetic and genetic silencing of CHFR in esophageal adenocarcinomas
- PMID: 20564104
- PMCID: PMC2930066
- DOI: 10.1002/cncr.25151
Epigenetic and genetic silencing of CHFR in esophageal adenocarcinomas
Abstract
Background: The checkpoint with forkhead-associated domain and RING-finger domain (CHFR) is a mitotic checkpoint protein with tumor-suppressor functions. In this study, the authors investigated the epigenetic and genetic mechanisms that regulate CHFR expression in esophageal adenocarcinomas (EACs).
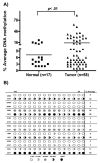
Methods: Quantitative reverse transcriptase polymerase chain reaction analysis demonstrated downregulation of CHFR transcript in 79% of EACs (44 of 56) compared with 41 normal samples (P < .001). Immunohistochemical analysis of CHFR protein expression showed absence or weak immunostaining for CHFR in 75% of EACs (56 of 75) compared with normal tissue samples. The authors next examined the promoter DNA hypermethylation of CHFR by using quantitative bisulfite pyrosequencing technology. They detected significant CHFR promoter DNA hypermethylation in 31% of tumor samples (18 of 58) compared with normal samples (P < .001). Treatment of OE33 cells with 5-Aza-deoxycytidine led to reduction in the promoter DNA methylation levels with restoration of the CHFR mRNA expression, which confirmed promoter DNA methylation as an epigenetic mechanism regulating CHFR expression. However, they identified several EACs where the CHFR mRNA expression was silenced in the absence of notable methylation. Therefore, the authors examined the relative DNA copy number level of CHFR compared with normal samples.
Results: The results confirmed a decrease or absence of the relative CHFR DNA copy number levels in 59% of tumor samples. Nine tumors that showed loss of CHFR mRNA expression, in absence of promoter DNA hypermethylation, demonstrated a significant loss of relative CHFR DNA copy numbers.
Conclusions: Taken together, their findings demonstrated that both epigenetic and genetic mechanisms were involved in silencing CHFR expression in EACs.
Cancer 2010. (c) 2010 American Cancer Society.
Figures




Similar articles
-
Emerging evidence for CHFR as a cancer biomarker: from tumor biology to precision medicine.Cancer Metastasis Rev. 2014 Mar;33(1):161-71. doi: 10.1007/s10555-013-9462-4. Cancer Metastasis Rev. 2014. PMID: 24375389 Free PMC article. Review.
-
CHFR: a key checkpoint component implicated in a wide range of cancers.Cell Mol Life Sci. 2012 May;69(10):1669-87. doi: 10.1007/s00018-011-0892-2. Epub 2011 Dec 13. Cell Mol Life Sci. 2012. PMID: 22159584 Free PMC article. Review.
-
Aberrant promoter hypermethylation of the CHFR gene in oral squamous cell carcinomas.Oncol Rep. 2009 Nov;22(5):1173-9. doi: 10.3892/or_00000552. Oncol Rep. 2009. PMID: 19787237
-
Promoter methylation of p16, Runx3, DAPK and CHFR genes is frequent in gastric carcinoma.Tumori. 2010 Sep-Oct;96(5):726-33. doi: 10.1177/030089161009600515. Tumori. 2010. PMID: 21302620
-
CHFR promoter hypermethylation and reduced CHFR mRNA expression in ovarian cancer.Int J Biol Markers. 2009 Apr-Jun;24(2):83-9. doi: 10.1177/172460080902400204. Int J Biol Markers. 2009. PMID: 19634111
Cited by
-
Genome-scale methylation analysis of Parkinson's disease patients' brains reveals DNA hypomethylation and increased mRNA expression of cytochrome P450 2E1.Neurogenetics. 2012 Feb;13(1):87-91. doi: 10.1007/s10048-011-0308-3. Epub 2012 Jan 12. Neurogenetics. 2012. PMID: 22238121
-
CHFR promotes the migration of human gastric cancer cells by inducing epithelial-to-mesenchymal transition in a HDAC1-dependent manner.Onco Targets Ther. 2019 Feb 7;12:1075-1084. doi: 10.2147/OTT.S191016. eCollection 2019. Onco Targets Ther. 2019. PMID: 30799937 Free PMC article.
-
Emerging evidence for CHFR as a cancer biomarker: from tumor biology to precision medicine.Cancer Metastasis Rev. 2014 Mar;33(1):161-71. doi: 10.1007/s10555-013-9462-4. Cancer Metastasis Rev. 2014. PMID: 24375389 Free PMC article. Review.
-
Multi-Omic Analysis of Esophageal Adenocarcinoma Uncovers Candidate Therapeutic Targets and Cancer-Selective Posttranscriptional Regulation.Mol Cell Proteomics. 2024 Jun;23(6):100764. doi: 10.1016/j.mcpro.2024.100764. Epub 2024 Apr 9. Mol Cell Proteomics. 2024. PMID: 38604503 Free PMC article.
-
CHFR: a key checkpoint component implicated in a wide range of cancers.Cell Mol Life Sci. 2012 May;69(10):1669-87. doi: 10.1007/s00018-011-0892-2. Epub 2011 Dec 13. Cell Mol Life Sci. 2012. PMID: 22159584 Free PMC article. Review.
References
-
- Devesa SS, Blot WJ, Fraumeni JF., Jr Changing patterns in the incidence of esophageal and gastric carcinoma in the United States. Cancer. 1998;83(10):2049–53. - PubMed
-
- Jankowski JA, Wright NA, Meltzer SJ, Triadafilopoulos G, Geboes K, Casson AG, et al. Molecular evolution of the metaplasia-dysplasia-adenocarcinoma sequence in the esophagus. Am J Pathol. 1999;154(4):965–73. Available from http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dop.... - PMC - PubMed
-
- Veugelers PJ, Porter GA, Guernsey DL, Casson AG. Obesity and lifestyle risk factors for gastroesophageal reflux disease, Barrett esophagus and esophageal adenocarcinoma. Dis Esophagus. 2006;19(5):321–8. Available from http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dop.... - PubMed
-
- Barrett MT, Sanchez CA, Prevo LJ, Wong DJ, Galipeau PC, Paulson TG, et al. Evolution of neoplastic cell lineages in Barrett oesophagus. Nat Genet. 1999;22(1):106–9. Available from http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dop.... - PMC - PubMed
-
- Zhuang Z, Vortmeyer AO, Mark EJ, Odze R, Emmert-Buck MR, Merino MJ, et al. Barrett's esophagus: metaplastic cells with loss of heterozygosity at the APC gene locus are clonal precursors to invasive adenocarcinoma. Cancer Res. 1996;56(9):1961–4. Available from http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dop.... - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

