The methylated N-terminal tail of RCC1 is required for stabilisation of its interaction with chromatin by Ran in live cells
- PMID: 20565941
- PMCID: PMC2898669
- DOI: 10.1186/1471-2121-11-43
The methylated N-terminal tail of RCC1 is required for stabilisation of its interaction with chromatin by Ran in live cells
Abstract
Background: Regulator of chromosome condensation 1 (RCC1) is the guanine nucleotide exchange factor for Ran GTPase. Localised generation of Ran-GTP by RCC1 on chromatin is critical for nucleocytoplasmic transport, mitotic spindle assembly and nuclear envelope formation. Both the N-terminal tail of RCC1 and its association with Ran are important for its interaction with chromatin in cells. In vitro, the association of Ran with RCC1 induces a conformational change in the N-terminal tail that promotes its interaction with DNA.
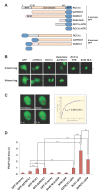
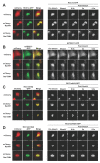
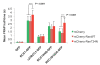
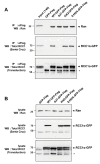
Results: We have investigated the mechanism of the dynamic interaction of the alpha isoform of human RCC1 (RCC1alpha) with chromatin in live cells using fluorescence recovery after photobleaching (FRAP) of green fluorescent protein (GFP) fusions. We show that the N-terminal tail stabilises the interaction of RCC1alpha with chromatin and this function can be partially replaced by another lysine-rich nuclear localisation signal. Removal of the tail prevents the interaction of RCC1alpha with chromatin from being stabilised by RanT24N, a mutant that binds stably to RCC1alpha. The interaction of RCC1alpha with chromatin is destabilised by mutation of lysine 4 (K4Q), which abolishes alpha-N-terminal methylation, and this interaction is no longer stabilised by RanT24N. However, alpha-N-terminal methylation of RCC1alpha is not regulated by the binding of RanT24N. Conversely, the association of Ran with precipitated RCC1alpha does not require the N-terminal tail of RCC1alpha or its methylation. The mobility of RCC1alpha on chromatin is increased by mutation of aspartate 182 (D182A), which inhibits guanine-nucleotide exchange activity, but RCC1alphaD182A can still bind nucleotide-free Ran and its interaction with chromatin is stabilised by RanT24N.
Conclusions: These results show that the stabilisation of the dynamic interaction of RCC1alpha with chromatin by Ran in live cells requires the N-terminal tail of RCC1alpha. alpha-N-methylation is not regulated by formation of the binary complex with Ran, but it promotes chromatin binding through the tail. This work supports a model in which the association of RCC1alpha with chromatin is promoted by a conformational change in the alpha-N-terminal methylated tail that is induced allosterically in the binary complex with Ran.
Figures

Similar articles
-
N-terminal alpha-methylation of RCC1 is necessary for stable chromatin association and normal mitosis.Nat Cell Biol. 2007 May;9(5):596-603. doi: 10.1038/ncb1572. Epub 2007 Apr 15. Nat Cell Biol. 2007. PMID: 17435751 Free PMC article.
-
RCC1 isoforms differ in their affinity for chromatin, molecular interactions and regulation by phosphorylation.J Cell Sci. 2007 Oct 1;120(Pt 19):3436-45. doi: 10.1242/jcs.009092. Epub 2007 Sep 12. J Cell Sci. 2007. PMID: 17855385
-
Regulation of chromatin binding by a conformational switch in the tail of the Ran exchange factor RCC1.J Cell Biol. 2008 Sep 8;182(5):827-36. doi: 10.1083/jcb.200803110. Epub 2008 Sep 1. J Cell Biol. 2008. PMID: 18762580 Free PMC article.
-
Premature chromatin condensation caused by loss of RCC1.Prog Cell Cycle Res. 2000;4:145-56. doi: 10.1007/978-1-4615-4253-7_13. Prog Cell Cycle Res. 2000. PMID: 10740822 Review.
-
The intricate roles of RCC1 in normal cells and cancer cells.Biochem Soc Trans. 2022 Feb 28;50(1):83-93. doi: 10.1042/BST20210861. Biochem Soc Trans. 2022. PMID: 35191966 Review.
Cited by
-
Past, present, and perspectives of protein N-terminal methylation.Curr Opin Chem Biol. 2021 Aug;63:115-122. doi: 10.1016/j.cbpa.2021.02.017. Epub 2021 Apr 8. Curr Opin Chem Biol. 2021. PMID: 33839647 Free PMC article. Review.
-
Importin-β modulates the permeability of the nuclear pore complex in a Ran-dependent manner.Elife. 2015 Mar 6;4:e04052. doi: 10.7554/eLife.04052. Elife. 2015. PMID: 25748139 Free PMC article.
-
The Multifaceted Roles of RCC1 in Tumorigenesis.Front Mol Biosci. 2020 Sep 15;7:225. doi: 10.3389/fmolb.2020.00225. eCollection 2020. Front Mol Biosci. 2020. PMID: 33102517 Free PMC article. Review.
-
Chromosomal gain promotes formation of a steep RanGTP gradient that drives mitosis in aneuploid cells.J Cell Biol. 2013 Jan 21;200(2):151-61. doi: 10.1083/jcb.201206142. Epub 2013 Jan 14. J Cell Biol. 2013. PMID: 23319601 Free PMC article.
-
Revisiting chromatin binding of the Arabidopsis UV-B photoreceptor UVR8.BMC Plant Biol. 2016 Feb 11;16:42. doi: 10.1186/s12870-016-0732-5. BMC Plant Biol. 2016. PMID: 26864020 Free PMC article.
References
-
- Ohtsubo M, Kai R, Furuno N, Sekiguchi T, Sekiguchi M, Hayashida H, Kuma K, Miyata T, Fukushige S, Murotsu T. Isolation and characterization of the active cDNA of the human cell cycle gene (RCC1) involved in the regulation of onset of chromosome condensation. Genes Dev. 1987;1:585–593. doi: 10.1101/gad.1.6.585. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

