Small RNA profiling reveals antisense transcription throughout the KSHV genome and novel small RNAs
- PMID: 20566670
- PMCID: PMC2905754
- DOI: 10.1261/rna.1967910
Small RNA profiling reveals antisense transcription throughout the KSHV genome and novel small RNAs
Abstract
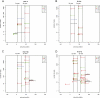
Kaposi's sarcoma-associated herpesvirus (KSHV) is a human tumor virus that encodes 12 precursor microRNAs (pre-miRNAs) that give rise to 17 different known approximately 22-nucleotide (nt) effector miRNAs. Like all herpesviruses, KSHV has two modes of infection: (1) a latent mode whereby only a subset of viral genes are expressed and (2) a lytic mode during which the full remaining viral genes are expressed. To date, KSHV miRNAs have been mostly identified via analysis of cells that are undergoing latent infection. Here, we developed a method to profile small RNAs ( approximately 18-75 nt) from populations of cells undergoing predominantly lytic infection. Using two different next-generation sequencing platforms, we cloned and sequenced both pre-miRNAs and derivative miRNAs. Our analysis shows that the vast majority of viral and host 5p miRNAs are co-terminal with the 5' end of the cloned pre-miRNAs, consistent with both being defined by microprocessor cleavage. We report the complete repertoire (25 total) of 5p and 3p derivative miRNAs from all 12 previously described KSHV pre-miRNAs. Two KSHV pre-miRNAs, pre-miR-K12-8 and pre-miR-K12-12, encode abundant derivative miRNAs from the previously unreported strands of the pre-miRNA. We identify several novel small RNAs of low abundance, including viral miRNA-offset-RNAs (moRNAs), and antisense viral miRNAs (miRNA-AS) that are encoded antisense to previously reported KSHV pre-miRNAs. Finally, we observe widespread antisense transcription relative to known coding sequences during lytic replication. Despite the enormous potential to form double-stranded RNA in KSHV-infected cells, we observe no evidence for the existence of abundant viral-derived small interfering RNAs (siRNAs).
Figures



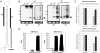


Similar articles
-
Kaposi's sarcoma-associated herpesvirus microRNAs repress breakpoint cluster region protein expression, enhance Rac1 activity, and increase in vitro angiogenesis.J Virol. 2015 Apr;89(8):4249-61. doi: 10.1128/JVI.03687-14. Epub 2015 Jan 28. J Virol. 2015. PMID: 25631082 Free PMC article.
-
Kaposi's sarcoma-associated herpesvirus expresses an array of viral microRNAs in latently infected cells.Proc Natl Acad Sci U S A. 2005 Apr 12;102(15):5570-5. doi: 10.1073/pnas.0408192102. Epub 2005 Mar 30. Proc Natl Acad Sci U S A. 2005. PMID: 15800047 Free PMC article.
-
Epigenetic regulation of Kaposi's sarcoma-associated herpesvirus latency by virus-encoded microRNAs that target Rta and the cellular Rbl2-DNMT pathway.J Virol. 2010 Mar;84(6):2697-706. doi: 10.1128/JVI.01997-09. Epub 2010 Jan 13. J Virol. 2010. PMID: 20071580 Free PMC article.
-
miRNAs and their roles in KSHV pathogenesis.Virus Res. 2019 Jun;266:15-24. doi: 10.1016/j.virusres.2019.03.024. Epub 2019 Apr 2. Virus Res. 2019. PMID: 30951791 Review.
-
Regulation of the MIR155 host gene in physiological and pathological processes.Gene. 2013 Dec 10;532(1):1-12. doi: 10.1016/j.gene.2012.12.009. Epub 2012 Dec 14. Gene. 2013. PMID: 23246696 Review.
Cited by
-
Looking at Kaposi's Sarcoma-Associated Herpesvirus-Host Interactions from a microRNA Viewpoint.Front Microbiol. 2012 Jan 11;2:271. doi: 10.3389/fmicb.2011.00271. eCollection 2011. Front Microbiol. 2012. PMID: 22275910 Free PMC article.
-
Kaposi's Sarcoma-Associated Herpesvirus microRNAs.Front Microbiol. 2012 May 3;3:165. doi: 10.3389/fmicb.2012.00165. eCollection 2012. Front Microbiol. 2012. PMID: 22563327 Free PMC article.
-
Insights into Polyomaviridae microRNA function derived from study of the bandicoot papillomatosis carcinomatosis viruses.J Virol. 2011 May;85(9):4487-500. doi: 10.1128/JVI.02557-10. Epub 2011 Feb 23. J Virol. 2011. PMID: 21345962 Free PMC article.
-
MicroRNA-mediated transformation by the Kaposi's sarcoma-associated herpesvirus Kaposin locus.J Virol. 2015 Feb;89(4):2333-41. doi: 10.1128/JVI.03317-14. Epub 2014 Dec 10. J Virol. 2015. PMID: 25505059 Free PMC article.
-
The viral and cellular microRNA targetome in lymphoblastoid cell lines.PLoS Pathog. 2012 Jan;8(1):e1002484. doi: 10.1371/journal.ppat.1002484. Epub 2012 Jan 26. PLoS Pathog. 2012. PMID: 22291592 Free PMC article.
References
-
- Applied Biosystems 2009. Whole-genome discovery and profiling of small RNAs using the SOLiD System. BioTechniques 46: 232–234
-
- Areste C, Blackbourn DJ 2009. Modulation of the immune system by Kaposi's sarcoma-associated herpesvirus. Trends Microbiol 17: 119–129 - PubMed
-
- Bartel DP 2004. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 116: 281–297 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
