Virion-associated cofactor high-mobility group DNA-binding protein-1 facilitates transposition from the herpes simplex virus/Sleeping Beauty amplicon vector platform
- PMID: 20568967
- PMCID: PMC2978546
- DOI: 10.1089/hum.2010.022
Virion-associated cofactor high-mobility group DNA-binding protein-1 facilitates transposition from the herpes simplex virus/Sleeping Beauty amplicon vector platform
Abstract
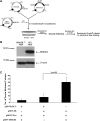
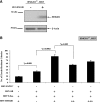
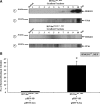
The development of the integration-competent, herpes simplex virus/Sleeping Beauty (HSV/SB) amplicon vector platform has created a means to efficiently and stably deliver therapeutic transcription units (termed "transgenons") to neurons within the mammalian brain. Furthermore, an investigation into the transposition capacity of the HSV/SB vector system revealed that the amplicon genome provides an optimal substrate for the transposition of transgenons at least 12 kb in length [de Silva, S., Mastrangelo, M.A., Lotta, L.T., Jr., Burris, C.A., Federoff, H.J., and Bowers, W.J. ( 2010 ). Gene Ther. 17, 424-431]. These results prompted an investigation into the factors that may contribute toward efficient transposition from the HSV/SB amplicon. One of the cellular cofactors known to play a key role during SB-mediated transposition is the high-mobility group DNA-binding protein-1 (HMGB1). Our present investigation into the role of HMGB1 during amplicon-based transposition revealed that transposition is not strictly dependent on the presence of cellular HMGB1, contrary to what had been previously demonstrated with plasmid-based SB transposition. We have shown for the first time that during amplicon preparation, biologically active HMGB1 derived from the packaging cell line is copackaged into amplicon vector particles. As a result, HSV/SB amplicon virions arrive prearmed with HMGB1 protein at levels sufficient for facilitating SB-mediated transposition in the transduced mammalian cell.
Figures

Similar articles
-
Targeting the central nervous system with herpes simplex virus / Sleeping Beauty hybrid amplicon vectors.Curr Gene Ther. 2011 Oct;11(5):332-40. doi: 10.2174/156652311797415845. Curr Gene Ther. 2011. PMID: 21711226 Free PMC article. Review.
-
Extending the transposable payload limit of Sleeping Beauty (SB) using the Herpes Simplex Virus (HSV)/SB amplicon-vector platform.Gene Ther. 2010 Mar;17(3):424-31. doi: 10.1038/gt.2009.144. Epub 2009 Oct 29. Gene Ther. 2010. PMID: 19865178 Free PMC article.
-
Herpes simplex virus/Sleeping Beauty vector-based embryonic gene transfer using the HSB5 mutant: loss of apparent transposition hyperactivity in vivo.Hum Gene Ther. 2010 Nov;21(11):1603-13. doi: 10.1089/hum.2010.062. Epub 2010 Oct 22. Hum Gene Ther. 2010. PMID: 20507234 Free PMC article.
-
The DNA-bending protein HMGB1 is a cellular cofactor of Sleeping Beauty transposition.Nucleic Acids Res. 2003 May 1;31(9):2313-22. doi: 10.1093/nar/gkg341. Nucleic Acids Res. 2003. PMID: 12711676 Free PMC article.
-
Minicircle-Based Engineering of Chimeric Antigen Receptor (CAR) T Cells.Recent Results Cancer Res. 2016;209:37-50. doi: 10.1007/978-3-319-42934-2_3. Recent Results Cancer Res. 2016. PMID: 28101686 Review.
Cited by
-
HMGB1 in health and disease.Mol Aspects Med. 2014 Dec;40:1-116. doi: 10.1016/j.mam.2014.05.001. Epub 2014 Jul 8. Mol Aspects Med. 2014. PMID: 25010388 Free PMC article. Review.
-
The Sleeping Beauty transposon system: a non-viral vector for gene therapy.Hum Mol Genet. 2011 Apr 15;20(R1):R14-20. doi: 10.1093/hmg/ddr140. Epub 2011 Apr 1. Hum Mol Genet. 2011. PMID: 21459777 Free PMC article. Review.
-
Gene transfer to the outflow tract.Exp Eye Res. 2017 May;158:73-84. doi: 10.1016/j.exer.2016.04.023. Epub 2016 Apr 27. Exp Eye Res. 2017. PMID: 27131906 Free PMC article. Review.
-
Targeting the central nervous system with herpes simplex virus / Sleeping Beauty hybrid amplicon vectors.Curr Gene Ther. 2011 Oct;11(5):332-40. doi: 10.2174/156652311797415845. Curr Gene Ther. 2011. PMID: 21711226 Free PMC article. Review.
-
β-globin matrix attachment region improves stable genomic expression of the Sleeping Beauty transposon.J Cell Biochem. 2011 Sep;112(9):2361-2375. doi: 10.1002/jcb.23159. J Cell Biochem. 2011. PMID: 21520245 Free PMC article.
References
-
- Bakowska J.C. Di Maria M.V. Camp S.M. Wang Y. Allen P.D. Breakefield X.O. Targeted transgene integration into transgenic mouse fibroblasts carrying the full-length human AAVS1 locus mediated by HSV/AAV rep+ hybrid amplicon vector. Gene Ther. 2003;10:1691–1702. - PubMed
-
- Bowers W.J. Federoff H.J. Herpes simplex virus type 1-derived amplicon vectors. In Gene Transfer: Delivery and Expression of DNA and RNA, A Laboratory Manual. In: Rossi J., editor; Friedmann T., editor. Cold Spring Harbor Press; Cold Spring Harbor, NY: 2006. pp. 227–254.
-
- Bowers W.J. Howard D.F. Brooks A.I. Halterman M.W. Federoff H.J. Expression of vhs and VP16 during HSV-1 helper virus-free amplicon packaging enhances titers. Gene Ther. 2001;8:111–120. - PubMed
-
- Bowers W.J. Mastrangelo M.A. Howard D.F. Southerland H.A. Maguire-Zeiss K.A. Federoff H.J. Neuronal precursor-restricted transduction via in utero CNS gene delivery of a novel bipartite HSV amplicon/transposase hybrid vector. Mol. Ther. 2006;13:580–588. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

