A draft physical map of a D-genome cotton species (Gossypium raimondii)
- PMID: 20569427
- PMCID: PMC2996926
- DOI: 10.1186/1471-2164-11-395
A draft physical map of a D-genome cotton species (Gossypium raimondii)
Abstract
Background: Genetically anchored physical maps of large eukaryotic genomes have proven useful both for their intrinsic merit and as an adjunct to genome sequencing. Cultivated tetraploid cottons, Gossypium hirsutum and G. barbadense, share a common ancestor formed by a merger of the A and D genomes about 1-2 million years ago. Toward the long-term goal of characterizing the spectrum of diversity among cotton genomes, the worldwide cotton community has prioritized the D genome progenitor Gossypium raimondii for complete sequencing.
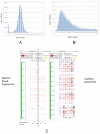
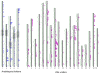
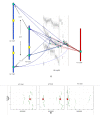
Results: A whole genome physical map of G. raimondii, the putative D genome ancestral species of tetraploid cottons was assembled, integrating genetically-anchored overgo hybridization probes, agarose based fingerprints and 'high information content fingerprinting' (HICF). A total of 13,662 BAC-end sequences and 2,828 DNA probes were used in genetically anchoring 1585 contigs to a cotton consensus genetic map, and 370 and 438 contigs, respectively to Arabidopsis thaliana (AT) and Vitis vinifera (VV) whole genome sequences.
Conclusion: Several lines of evidence suggest that the G. raimondii genome is comprised of two qualitatively different components. Much of the gene rich component is aligned to the Arabidopsis and Vitis vinifera genomes and shows promise for utilizing translational genomic approaches in understanding this important genome and its resident genes. The integrated genetic-physical map is of value both in assembling and validating a planned reference sequence.
Figures

References
-
- Wendel JF, Albert VA. Phylogenetics of the Cotton genus (Gossypium): Character-state weighted parsimony analysis of chloroplast-DNA restriction site data and its systematic and biogeographic implications. Systematic Botany. 1992;17:115–143. doi: 10.2307/2419069. - DOI
-
- Rong J, Abbey C, Bowers JE, Brubaker CL, Chang C, Chee PW, Delmonte TA, Ding X, Garza JJ, Marler BS. et al. A 3347-locus genetic recombination map of sequence-tagged sites reveals features of genome organization, transmission and evolution of cotton (Gossypium) Genetics. 2004;166:389–417. doi: 10.1534/genetics.166.1.389. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

