A distinct spectrum of copy number aberrations in pediatric high-grade gliomas
- PMID: 20570930
- PMCID: PMC2896553
- DOI: 10.1158/1078-0432.CCR-10-0438
A distinct spectrum of copy number aberrations in pediatric high-grade gliomas
Abstract
Purpose: As genome-scale technologies begin to unravel the complexity of the equivalent tumors in adults, we can attempt detailed characterization of high-grade gliomas in children, that have until recently been lacking. Toward this end, we sought to validate and extend investigations of the differences between pediatric and adult tumors.
Experimental design: We carried out copy number profiling by array comparative genomic hybridization using a 32K bacterial artificial chromosome platform on 63 formalin-fixed paraffin-embedded cases of high-grade glioma arising in children and young people (<23 years).
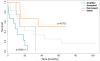
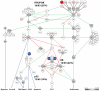
Results: The genomic profiles of these tumors could be subclassified into four categories: those with stable genomes, which were associated with a better prognosis; those with aneuploid and those with highly rearranged genomes; and those with an amplifier genotype, which had a significantly worse clinical outcome. Independent of this was a clear segregation of cases with 1q gain (more common in children) from those with concurrent 7 gain/10q loss (a defining feature of adults). Detailed mapping of all the amplification and deletion events revealed numerous low-frequency amplifications, including IGF1R, PDGFRB, PIK3CA, CDK6, CCND1, and CCNE1, and novel homozygous deletions encompassing unknown genes, including those at 5q35, 10q25, and 22q13. Despite this, aberrations targeting the "core signaling pathways" in adult glioblastomas are significantly underrepresented in the pediatric setting.
Conclusions: These data highlight that although there are overlaps in the genomic events driving gliomagenesis of all ages, the pediatric disease harbors a distinct spectrum of copy number aberrations compared with adults.
Figures



References
-
- Rao SK, Edwards J, Joshi AD, Siu IM, Riggins GJ. A survey of glioblastoma genomic amplifications and deletions. J Neurooncol. 2009 - PubMed
-
- de Tayrac M, Etcheverry A, Aubry M, et al. Integrative genome-wide analysis reveals a robust genomic glioblastoma signature associated with copy number driving changes in gene expression. Genes Chromosomes Cancer. 2009;48:55–68. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

