Isolated short CTG/CAG DNA slip-outs are repaired efficiently by hMutSbeta, but clustered slip-outs are poorly repaired
- PMID: 20571119
- PMCID: PMC2906547
- DOI: 10.1073/pnas.0909087107
Isolated short CTG/CAG DNA slip-outs are repaired efficiently by hMutSbeta, but clustered slip-outs are poorly repaired
Abstract
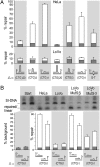
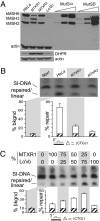
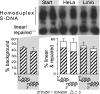
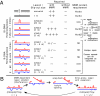
Expansions of CTG/CAG trinucleotide repeats, thought to involve slipped DNAs at the repeats, cause numerous diseases including myotonic dystrophy and Huntington's disease. By unknown mechanisms, further repeat expansions in transgenic mice carrying expanded CTG/CAG tracts require the mismatch repair (MMR) proteins MSH2 and MSH3, forming the MutSbeta complex. Using an in vitro repair assay, we investigated the effect of slip-out size, with lengths of 1, 3, or 20 excess CTG repeats, as well as the effect of the number of slip-outs per molecule, on the requirement for human MMR. Long slip-outs escaped repair, whereas short slip-outs were repaired efficiently, much greater than a G-T mismatch, but required hMutSbeta. Higher or lower levels of hMutSbeta or its complete absence were detrimental to proper repair of short slip-outs. Surprisingly, clusters of as many as 62 short slip-outs (one to three repeat units each) along a single DNA molecule with (CTG)50*(CAG)50 repeats were refractory to repair, and repair efficiency was reduced further without MMR. Consistent with the MutSbeta requirement for instability, hMutSbeta is required to process isolated short slip-outs; however, multiple adjacent short slip-outs block each other's repair, possibly acting as roadblocks to progression of repair and allowing error-prone repair. Results suggest that expansions can arise by escaped repair of long slip-outs, tandem short slip-outs, or isolated short slip-outs; the latter two types are sensitive to hMutSbeta. Poor repair of clustered DNA lesions has previously been associated only with ionizing radiation damage. Our results extend this interference in repair to neurodegenerative disease-causing mutations in which clustered slip-outs escape proper repair and lead to expansions.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
Human mismatch repair protein hMutLα is required to repair short slipped-DNAs of trinucleotide repeats.J Biol Chem. 2012 Dec 7;287(50):41844-50. doi: 10.1074/jbc.M112.420398. Epub 2012 Oct 18. J Biol Chem. 2012. PMID: 23086927 Free PMC article.
-
Absence of MutSβ leads to the formation of slipped-DNA for CTG/CAG contractions at primate replication forks.DNA Repair (Amst). 2016 Jun;42:107-18. doi: 10.1016/j.dnarep.2016.04.002. Epub 2016 Apr 16. DNA Repair (Amst). 2016. PMID: 27155933 Free PMC article.
-
MSH2 ATPase domain mutation affects CTG*CAG repeat instability in transgenic mice.PLoS Genet. 2009 May;5(5):e1000482. doi: 10.1371/journal.pgen.1000482. Epub 2009 May 15. PLoS Genet. 2009. PMID: 19436705 Free PMC article.
-
Disease-associated repeat instability and mismatch repair.DNA Repair (Amst). 2016 Feb;38:117-126. doi: 10.1016/j.dnarep.2015.11.008. Epub 2015 Dec 12. DNA Repair (Amst). 2016. PMID: 26774442 Review.
-
DNA mismatch repair in trinucleotide repeat instability.Sci China Life Sci. 2017 Oct;60(10):1087-1092. doi: 10.1007/s11427-017-9186-7. Epub 2017 Oct 24. Sci China Life Sci. 2017. PMID: 29075942 Free PMC article. Review.
Cited by
-
Triptycene-based small molecules modulate (CAG)·(CTG) repeat junctions.Chem Sci. 2015 Aug 14;6(8):4752-4755. doi: 10.1039/c5sc01595b. Epub 2015 Jun 10. Chem Sci. 2015. PMID: 26366282 Free PMC article.
-
Trinucleotide repeat expansions catalyzed by human cell-free extracts.Cell Res. 2013 Apr;23(4):565-72. doi: 10.1038/cr.2013.12. Epub 2013 Jan 22. Cell Res. 2013. PMID: 23337586 Free PMC article.
-
Tissue- and age-specific DNA replication patterns at the CTG/CAG-expanded human myotonic dystrophy type 1 locus.Nat Struct Mol Biol. 2010 Sep;17(9):1079-87. doi: 10.1038/nsmb.1876. Epub 2010 Aug 15. Nat Struct Mol Biol. 2010. PMID: 20711191
-
The G-rich Repeats in FMR1 and C9orf72 Loci Are Hotspots for Local Unpairing of DNA.Genetics. 2018 Dec;210(4):1239-1252. doi: 10.1534/genetics.118.301672. Epub 2018 Nov 5. Genetics. 2018. PMID: 30396881 Free PMC article.
-
Sequence length dictates repeated CAG folding in three-way junctions.Biochemistry. 2011 Feb 1;50(4):458-65. doi: 10.1021/bi101756e. Epub 2010 Dec 31. Biochemistry. 2011. PMID: 21142085 Free PMC article.
References
-
- Pearson CE, Nichol Edamura K, Cleary JD. Repeat instability: Mechanisms of dynamic mutations. Nat Rev Genet. 2005;6:729–742. - PubMed
-
- López Castel A, Cleary JD, Pearson CE. Repeat instability as the basis for human diseases and as a potential target for therapy. Nat Rev Mol Cell Biol. 2010;11:165–170. - PubMed
-
- Slean MM, Panigrahi GB, Ranum LP, Pearson CE. Mutagenic roles of DNA “repair” proteins in antibody diversity and disease-associated trinucleotide repeat instability. DNA Repair (Amst) 2008;7:1135–1154. - PubMed
-
- Li GM. Mechanisms and functions of DNA mismatch repair. Cell Res. 2008;18:85–98. - PubMed
-
- Littman SJ, Fang WH, Modrich P. Repair of large insertion/deletion heterologies in human nuclear extracts is directed by a 5′ single-strand break and is independent of the mismatch repair system. J Biol Chem. 1999;274:7474–7481. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

