DPARSF: A MATLAB Toolbox for "Pipeline" Data Analysis of Resting-State fMRI
- PMID: 20577591
- PMCID: PMC2889691
- DOI: 10.3389/fnsys.2010.00013
DPARSF: A MATLAB Toolbox for "Pipeline" Data Analysis of Resting-State fMRI
Abstract
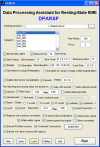
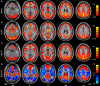
Resting-state functional magnetic resonance imaging (fMRI) has attracted more and more attention because of its effectiveness, simplicity and non-invasiveness in exploration of the intrinsic functional architecture of the human brain. However, user-friendly toolbox for "pipeline" data analysis of resting-state fMRI is still lacking. Based on some functions in Statistical Parametric Mapping (SPM) and Resting-State fMRI Data Analysis Toolkit (REST), we have developed a MATLAB toolbox called Data Processing Assistant for Resting-State fMRI (DPARSF) for "pipeline" data analysis of resting-state fMRI. After the user arranges the Digital Imaging and Communications in Medicine (DICOM) files and click a few buttons to set parameters, DPARSF will then give all the preprocessed (slice timing, realign, normalize, smooth) data and results for functional connectivity, regional homogeneity, amplitude of low-frequency fluctuation (ALFF), and fractional ALFF. DPARSF can also create a report for excluding subjects with excessive head motion and generate a set of pictures for easily checking the effect of normalization. In addition, users can also use DPARSF to extract time courses from regions of interest.
Keywords: DPARSF; REST; SPM; data analysis; resting-state fMRI.
Figures

References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

