A viral microRNA down-regulates multiple cell cycle genes through mRNA 5'UTRs
- PMID: 20585629
- PMCID: PMC2891821
- DOI: 10.1371/journal.ppat.1000967
A viral microRNA down-regulates multiple cell cycle genes through mRNA 5'UTRs
Abstract
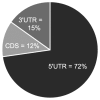
Global gene expression data combined with bioinformatic analysis provides strong evidence that mammalian miRNAs mediate repression of gene expression primarily through binding sites within the 3' untranslated region (UTR). Using RNA induced silencing complex immunoprecipitation (RISC-IP) techniques we have identified multiple cellular targets for a human cytomegalovirus (HCMV) miRNA, miR-US25-1. Strikingly, this miRNA binds target sites primarily within 5'UTRs, mediating significant reduction in gene expression. Intriguingly, many of the genes targeted by miR-US25-1 are associated with cell cycle control, including cyclin E2, BRCC3, EID1, MAPRE2, and CD147, suggesting that miR-US25-1 is targeting genes within a related pathway. Deletion of miR-US25-1 from HCMV results in over expression of cyclin E2 in the context of viral infection. Our studies demonstrate that a viral miRNA mediates translational repression of multiple cellular genes by targeting mRNA 5'UTRs.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
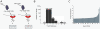
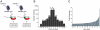


References
-
- Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–297. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

