BioModels Database: An enhanced, curated and annotated resource for published quantitative kinetic models
- PMID: 20587024
- PMCID: PMC2909940
- DOI: 10.1186/1752-0509-4-92
BioModels Database: An enhanced, curated and annotated resource for published quantitative kinetic models
Abstract
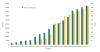
Background: Quantitative models of biochemical and cellular systems are used to answer a variety of questions in the biological sciences. The number of published quantitative models is growing steadily thanks to increasing interest in the use of models as well as the development of improved software systems and the availability of better, cheaper computer hardware. To maximise the benefits of this growing body of models, the field needs centralised model repositories that will encourage, facilitate and promote model dissemination and reuse. Ideally, the models stored in these repositories should be extensively tested and encoded in community-supported and standardised formats. In addition, the models and their components should be cross-referenced with other resources in order to allow their unambiguous identification.
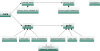
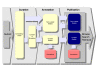
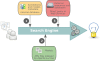
Description: BioModels Database http://www.ebi.ac.uk/biomodels/ is aimed at addressing exactly these needs. It is a freely-accessible online resource for storing, viewing, retrieving, and analysing published, peer-reviewed quantitative models of biochemical and cellular systems. The structure and behaviour of each simulation model distributed by BioModels Database are thoroughly checked; in addition, model elements are annotated with terms from controlled vocabularies as well as linked to relevant data resources. Models can be examined online or downloaded in various formats. Reaction network diagrams generated from the models are also available in several formats. BioModels Database also provides features such as online simulation and the extraction of components from large scale models into smaller submodels. Finally, the system provides a range of web services that external software systems can use to access up-to-date data from the database.
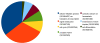
Conclusions: BioModels Database has become a recognised reference resource for systems biology. It is being used by the community in a variety of ways; for example, it is used to benchmark different simulation systems, and to study the clustering of models based upon their annotations. Model deposition to the database today is advised by several publishers of scientific journals. The models in BioModels Database are freely distributed and reusable; the underlying software infrastructure is also available from SourceForge https://sourceforge.net/projects/biomodels/ under the GNU General Public License.
Figures


References
-
- Hucka M, Finney A, Sauro HM, Bolouri H, Doyle JC, Kitano H, Arkin AP, Bornstein BJ, Bray D, Cornish-Bowden A, Cuellar AA, Dronov S, Gilles ED, Ginkel M, Gor V, Goryanin II, Hedley WJ, Hodgman TC, Hofmeyr JH, Hunter PJ, Juty NS, Kasberger JL, Kremling A, Kummer U, Le Novère N, Loew LM, Lucio D, Mendes P, Minch E, Mjolsness ED, Nakayama Y, Nelson MR, Nielsen PF, Sakurada T, Schaff JC, Shapiro BE, Shimizu TS, Spence HD, Stelling J, Takahashi K, Tomita M, Wagner J, Wang J. The systems biology markup language (SBML): a medium for representation and exchange of biochemical network models. Bioinformatics. 2003;19(4):524–531. doi: 10.1093/bioinformatics/btg015. - DOI - PubMed
-
- Systems Biology Markup Language (SBML) http://sbml.org/
-
- Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, Matese JC, Richardson JE, Ringwald M, Sherlock GMRG. Gene Ontology: tool for the unification of biology. Nature Genetics. 2000;25:25–29. doi: 10.1038/75556. - DOI - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

