The Rsr1/Bud1 GTPase interacts with itself and the Cdc42 GTPase during bud-site selection and polarity establishment in budding yeast
- PMID: 20587777
- PMCID: PMC2929994
- DOI: 10.1091/mbc.E10-03-0232
The Rsr1/Bud1 GTPase interacts with itself and the Cdc42 GTPase during bud-site selection and polarity establishment in budding yeast
Abstract
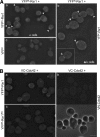
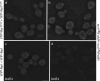
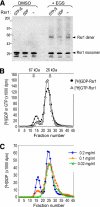
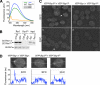
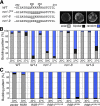
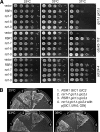
Cell polarization occurs along a single axis that is generally determined in response to spatial cues. In budding yeast, the Rsr1 GTPase and its regulators direct the establishment of cell polarity at the proper cortical location in response to cell type-specific cues. Here we use a combination of in vivo and in vitro approaches to understand how Rsr1 polarization is established. We find that Rsr1 associates with itself in a spatially and temporally controlled manner. The homotypic interaction and localization of Rsr1 to the mother-bud neck and to the subsequent division site are dependent on its GDP-GTP exchange factor Bud5. Analyses of rsr1 mutants suggest that Bud5 recruits Rsr1 to these sites and promotes the homodimer formation. Rsr1 also exhibits heterotypic interaction with the Cdc42 GTPase in vivo. We show that the polybasic region of Rsr1 is necessary for the efficient homotypic and heterotypic interactions, selection of a proper growth site, and polarity establishment. Our findings thus suggest that dimerization of GTPases may be an efficient mechanism to set up cellular asymmetry.
Figures

Similar articles
-
Temporal regulation of cell polarity via the interaction of the Ras GTPase Rsr1 and the scaffold protein Bem1.Mol Biol Cell. 2019 Sep 15;30(20):2543-2557. doi: 10.1091/mbc.E19-02-0106. Epub 2019 Aug 14. Mol Biol Cell. 2019. PMID: 31411940 Free PMC article.
-
Interaction between a Ras and a Rho GTPase couples selection of a growth site to the development of cell polarity in yeast.Mol Biol Cell. 2003 Dec;14(12):4958-70. doi: 10.1091/mbc.e03-06-0426. Epub 2003 Sep 5. Mol Biol Cell. 2003. PMID: 12960420 Free PMC article.
-
The upstream regulator, Rsr1p, and downstream effectors, Gic1p and Gic2p, of the Cdc42p small GTPase coordinately regulate initiation of budding in Saccharomyces cerevisiae.Genes Cells. 2003 Mar;8(3):235-50. doi: 10.1046/j.1365-2443.2003.00629.x. Genes Cells. 2003. PMID: 12622721
-
Cell polarity: connecting to the cortex.Curr Biol. 2001 Aug 7;11(15):R610-2. doi: 10.1016/s0960-9822(01)00365-7. Curr Biol. 2001. PMID: 11516968 Review.
-
Regulation of Cdc42 for polarized growth in budding yeast.Microb Cell. 2020 May 19;7(7):175-189. doi: 10.15698/mic2020.07.722. Microb Cell. 2020. PMID: 32656257 Free PMC article. Review.
Cited by
-
The shared role of the Rsr1 GTPase and Gic1/Gic2 in Cdc42 polarization.Mol Biol Cell. 2018 Oct 1;29(20):2359-2369. doi: 10.1091/mbc.E18-02-0145. Epub 2018 Aug 9. Mol Biol Cell. 2018. PMID: 30091649 Free PMC article.
-
Mating yeast cells use an intrinsic polarity site to assemble a pheromone-gradient tracking machine.J Cell Biol. 2019 Nov 4;218(11):3730-3752. doi: 10.1083/jcb.201901155. Epub 2019 Sep 30. J Cell Biol. 2019. PMID: 31570500 Free PMC article.
-
Computational and biochemical characterization of two partially overlapping interfaces and multiple weak-affinity K-Ras dimers.Sci Rep. 2017 Jan 9;7:40109. doi: 10.1038/srep40109. Sci Rep. 2017. PMID: 28067274 Free PMC article.
-
Two Rac paralogs regulate polarized growth in the human fungal pathogen Cryptococcus neoformans.Fungal Genet Biol. 2013 Aug;57:58-75. doi: 10.1016/j.fgb.2013.05.006. Epub 2013 Jun 5. Fungal Genet Biol. 2013. PMID: 23748012 Free PMC article.
-
Coupling of septins to the axial landmark by Bud4 in budding yeast.J Cell Sci. 2013 Mar 1;126(Pt 5):1218-26. doi: 10.1242/jcs.118521. Epub 2013 Jan 23. J Cell Sci. 2013. PMID: 23345395 Free PMC article.
References
-
- Ausubel F. M., Brent R., Kingston R. E., Moore D. D., Seidman J. G., Struhl K. New York: John Wiley & Sons; 1999. Current Protocols in Molecular Biology.
-
- Bailly E., Cabantous S., Sondaz D., Bernadac A., Simon M.-N. Differential cellular localization among mitotic cyclins from Saccharomyces cerevisiae: a new role for the axial budding protein Bud3 in targeting Clb2 to the mother-bud neck. J. Cell Sci. 2003;116:4119–4130. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

