Similarity of cytochrome c oxidases in different organisms
- PMID: 20589635
- PMCID: PMC4220736
- DOI: 10.1002/prot.22783
Similarity of cytochrome c oxidases in different organisms
Abstract
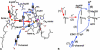
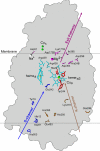
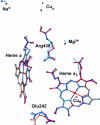
Most of biological oxygen reduction is catalyzed by the heme-copper oxygen reductases. These enzymes are redox-driven proton pumps that take part in generating the proton gradient in both prokaryotes and mitochondria that drives synthesis of ATP. The enzymes have been divided into three evolutionarily-related groups: the A-, B-, and C-families. Recent comparative studies suggest that all oxygen reductases perform the same chemistry for oxygen reduction and comprise the same essential elements of the proton pumping mechanism, such as the proton loading and kinetic gating sites, which, however, appear to be different in different families. All species of the A-family, however, demonstrate remarkable similarity of the central processing unit of the enzyme, as revealed by their recent crystal structures. Here we demonstrate that cytochrome c oxidases (CcO) of such diverse organisms as a mammal (bovine heart mitochondrial CcO), photosynthetic bacteria (Rhodobacter sphaeroides CcO), and soil bacteria (Paracoccus denitrificans CcO) are not only structurally similar, but almost identical in microscopic electrostatics and thermodynamics properties of their key amino-acids. By using pK(a) calculations of some of the key residues of the catalytic site, D- and K- proton input, and putative proton output channels of these three different enzymes, we demonstrate that the microscopic properties of key residues are almost identical, which strongly suggests the same mechanism in these species. The quantitative precision with which the microscopic physical properties of these enzymes have remained constant despite different evolutionary routes undertaken is striking.
Copyright 2010 Wiley-Liss, Inc.
Figures
Similar articles
-
Comparative genomics and site-directed mutagenesis support the existence of only one input channel for protons in the C-family (cbb3 oxidase) of heme-copper oxygen reductases.Biochemistry. 2007 Sep 4;46(35):9963-72. doi: 10.1021/bi700659y. Epub 2007 Aug 4. Biochemistry. 2007. PMID: 17676874
-
Characterizing the proton loading site in cytochrome c oxidase.Proc Natl Acad Sci U S A. 2014 Aug 26;111(34):12414-9. doi: 10.1073/pnas.1407187111. Epub 2014 Aug 11. Proc Natl Acad Sci U S A. 2014. PMID: 25114210 Free PMC article.
-
Mutations in the putative H-channel in the cytochrome c oxidase from Rhodobacter sphaeroides show that this channel is not important for proton conduction but reveal modulation of the properties of heme a.Biochemistry. 2000 Mar 21;39(11):2989-96. doi: 10.1021/bi9924821. Biochemistry. 2000. PMID: 10715119
-
Protein structure: proton-pumping oxidases.Curr Biol. 1996 Jan 1;6(1):36-8. doi: 10.1016/s0960-9822(02)00416-5. Curr Biol. 1996. PMID: 8825522 Review.
-
A cooperative model for proton pumping in cytochrome c oxidase.Biochim Biophys Acta. 2004 Apr 12;1655(1-3):353-64. doi: 10.1016/j.bbabio.2003.06.002. Biochim Biophys Acta. 2004. PMID: 15100051 Review.
Cited by
-
A common mechanism explains the induction of aerobic fermentation and adaptive antioxidant response in Phaffia rhodozyma.Microb Cell Fact. 2018 Apr 3;17(1):53. doi: 10.1186/s12934-018-0898-7. Microb Cell Fact. 2018. PMID: 29615045 Free PMC article.
-
The Redox-Active Tyrosine Is Essential for Proton Pumping in Cytochrome c Oxidase.Front Chem. 2021 Apr 14;9:640155. doi: 10.3389/fchem.2021.640155. eCollection 2021. Front Chem. 2021. PMID: 33937193 Free PMC article.
-
Control of transmembrane charge transfer in cytochrome c oxidase by the membrane potential.Nat Commun. 2018 Aug 9;9(1):3187. doi: 10.1038/s41467-018-05615-5. Nat Commun. 2018. PMID: 30093670 Free PMC article.
-
Deciphering the mechanism by which the yeast Phaffia rhodozyma responds adaptively to environmental, nutritional, and genetic cues.J Ind Microbiol Biotechnol. 2021 Dec 23;48(9-10):kuab048. doi: 10.1093/jimb/kuab048. J Ind Microbiol Biotechnol. 2021. PMID: 34302341 Free PMC article. Review.
-
Diversity of Cytochrome c Oxidase Assembly Proteins in Bacteria.Microorganisms. 2022 Apr 28;10(5):926. doi: 10.3390/microorganisms10050926. Microorganisms. 2022. PMID: 35630371 Free PMC article. Review.
References
-
- Gennis RB. Coupled proton and electron transfer reactions in cytochrome oxidase. Front Biosci. 2004;9:581–591. - PubMed
-
- Wikstrom M. Proton translocation by bacteriorhodopsin and heme-copper oxidases. Curr Opin Struct Biol. 1998;8:480–488. - PubMed
-
- Belevich I, Verkhovsky MI. Molecular mechanism of proton translocation by cytochrome c oxidase. Antioxidants & Redox Signaling. 2008;10(1):1–29. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

