Proteins that switch folds
- PMID: 20591649
- PMCID: PMC2928869
- DOI: 10.1016/j.sbi.2010.06.002
Proteins that switch folds
Abstract
An increasing number of proteins demonstrate the ability to switch between very different fold topologies, expanding their functional utility through new binding interactions. Recent examples of fold switching from naturally occurring and designed systems have a number of common features: (i) The structural transitions require states with diminished stability; (ii) Switching involves flexible regions in one conformer or the other; (iii) A new binding surface is revealed in the alternate fold that can lead to both stabilization of the alternative state and expansion of biological function. Fold switching not only provides insight into how new folds evolve, but also indicates that an amino acid sequence has more information content than previously thought. A polypeptide chain can encode a stable fold while simultaneously hiding latent propensities for alternative states with novel functions.
Copyright (c) 2010 Elsevier Ltd. All rights reserved.
Figures
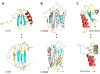
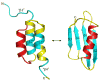
References
-
- Anfinsen CB. Principles that govern the folding of protein chains. Science. 1973;181:223–230. - PubMed
-
- Mottonen J, Strand A, Symersky J, Sweet RM, Danley DE, Geoghegan KF, Gerard RD, Goldsmith EJ. Structural basis of latency in plasminogen activator inhibitor-1. Nature. 1992;355:270–273. - PubMed
-
- Yamasaki M, Li W, Johnson DJ, Huntington JA. Crystal structure of a stable dimer reveals the molecular basis of serpin polymerization. Nature. 2008;455:1255–1258. - PubMed
-
- Bullough PA, Hughson FM, Skehel JJ, Wiley DC. Structure of influenza haemagglutinin at the pH of membrane fusion. Nature. 1994;371:37–43. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

