Class-specific restrictions define primase interactions with DNA template and replicative helicase
- PMID: 20591822
- PMCID: PMC2978363
- DOI: 10.1093/nar/gkq588
Class-specific restrictions define primase interactions with DNA template and replicative helicase
Abstract
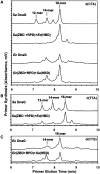
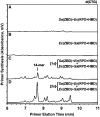
Bacterial primase is stimulated by replicative helicase to produce RNA primers that are essential for DNA replication. To identify mechanisms regulating primase activity, we characterized primase initiation specificity and interactions with the replicative helicase for gram-positive Firmicutes (Staphylococcus, Bacillus and Geobacillus) and gram-negative Proteobacteria (Escherichia, Yersinia and Pseudomonas). Contributions of the primase zinc-binding domain, RNA polymerase domain and helicase-binding domain on de novo primer synthesis were determined using mutated, truncated, chimeric and wild-type primases. Key residues in the β4 strand of the primase zinc-binding domain defined class-associated trinucleotide recognition and substitution of these amino acids transferred specificity across classes. A change in template recognition provided functional evidence for interaction in trans between the zinc-binding domain and RNA polymerase domain of two separate primases. Helicase binding to the primase C-terminal helicase-binding domain modulated RNA primer length in a species-specific manner and productive interactions paralleled genetic relatedness. Results demonstrated that primase template specificity is conserved within a bacterial class, whereas the primase-helicase interaction has co-evolved within each species.
Figures




Similar articles
-
Staphylococcus aureus helicase but not Escherichia coli helicase stimulates S. aureus primase activity and maintains initiation specificity.J Bacteriol. 2006 Jul;188(13):4673-80. doi: 10.1128/JB.00316-06. J Bacteriol. 2006. PMID: 16788176 Free PMC article.
-
Plant organellar DNA primase-helicase synthesizes RNA primers for organellar DNA polymerases using a unique recognition sequence.Nucleic Acids Res. 2017 Oct 13;45(18):10764-10774. doi: 10.1093/nar/gkx745. Nucleic Acids Res. 2017. PMID: 28977480 Free PMC article.
-
Structure of the zinc-binding domain of Bacillus stearothermophilus DNA primase.Structure. 2000 Mar 15;8(3):231-9. doi: 10.1016/s0969-2126(00)00101-5. Structure. 2000. PMID: 10745010
-
The bacterial helicase-primase interaction: a common structural/functional module.Structure. 2005 Jun;13(6):839-44. doi: 10.1016/j.str.2005.04.006. Structure. 2005. PMID: 15939015 Free PMC article. Review.
-
Regulation of bacterial priming and daughter strand synthesis through helicase-primase interactions.Nucleic Acids Res. 2006;34(15):4082-8. doi: 10.1093/nar/gkl363. Epub 2006 Aug 25. Nucleic Acids Res. 2006. PMID: 16935873 Free PMC article. Review.
Cited by
-
Unravelling staphylococcal small-colony variants in cardiac implantable electronic device infections: clinical characteristics, management, and genomic insights.Front Cell Infect Microbiol. 2024 Jan 8;13:1321626. doi: 10.3389/fcimb.2023.1321626. eCollection 2023. Front Cell Infect Microbiol. 2024. PMID: 38259974 Free PMC article.
-
Architecture and conservation of the bacterial DNA replication machinery, an underexploited drug target.Curr Drug Targets. 2012 Mar;13(3):352-72. doi: 10.2174/138945012799424598. Curr Drug Targets. 2012. PMID: 22206257 Free PMC article. Review.
-
Nucleic Acid Amplification Based Diagnostic of Lyme (Neuro-)borreliosis - Lost in the Jungle of Methods, Targets, and Assays?Open Neurol J. 2012;6:129-39. doi: 10.2174/1874205X01206010129. Epub 2012 Nov 16. Open Neurol J. 2012. PMID: 23230454 Free PMC article.
-
Structural Insight into the Specific DNA Template Binding to DnaG primase in Bacteria.Sci Rep. 2017 Apr 6;7(1):659. doi: 10.1038/s41598-017-00767-8. Sci Rep. 2017. PMID: 28386108 Free PMC article.
-
The Macromolecular Machines that Duplicate the Escherichia coli Chromosome as Targets for Drug Discovery.Antibiotics (Basel). 2018 Mar 14;7(1):23. doi: 10.3390/antibiotics7010023. Antibiotics (Basel). 2018. PMID: 29538288 Free PMC article. Review.
References
-
- Kornberg A, Baker T. DNA Replication. 2nd edn. Freeman: San Francisco; 1992.
-
- Wu CA, Zechner EL, Reems JA, McHenry CS, Marians KJ. Coordinated leading- and lagging-strand synthesis at the Escherichia coli DNA replication fork: V. Primase action regulates the cycle of Okazaki fragment synthesis. J. Biol. Chem. 1992;267:4074–4083. - PubMed
-
- Pan H, Wigley DB. Structure of the zinc-binding domain of Bacillus stearothermophilus DNA primase. Structure. 2000;8:231–239. - PubMed
-
- Griep MA, Lokey ER. The role of zinc and the reactivity of cysteines in Escherichia coli primase. Biochemistry. 1996;35:8260–8267. - PubMed

