Mapping surface accessibility of the C1r/C1s tetramer by chemical modification and mass spectrometry provides new insights into assembly of the human C1 complex
- PMID: 20592021
- PMCID: PMC2952226
- DOI: 10.1074/jbc.M110.149112
Mapping surface accessibility of the C1r/C1s tetramer by chemical modification and mass spectrometry provides new insights into assembly of the human C1 complex
Abstract
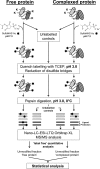
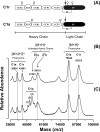
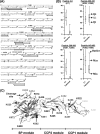
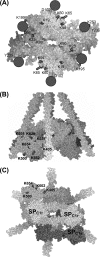
C1, the complex that triggers the classic pathway of complement, is a 790-kDa assembly resulting from association of a recognition protein C1q with a Ca(2+)-dependent tetramer comprising two copies of the proteases C1r and C1s. Early structural investigations have shown that the extended C1s-C1r-C1r-C1s tetramer folds into a compact conformation in C1. Recent site-directed mutagenesis studies have identified the C1q-binding sites in C1r and C1s and led to a three-dimensional model of the C1 complex (Bally, I., Rossi, V., Lunardi, T., Thielens, N. M., Gaboriaud, C., and Arlaud, G. J. (2009) J. Biol. Chem. 284, 19340-19348). In this study, we have used a mass spectrometry-based strategy involving a label-free semi-quantitative analysis of protein samples to gain new structural insights into C1 assembly. Using a stable chemical modification, we have compared the accessibility of the lysine residues in the isolated tetramer and in C1. The labeling data account for 51 of the 73 lysine residues of C1r and C1s. They strongly support the hypothesis that both C1s CUB(1)-EGF-CUB(2) interaction domains, which are distant in the free tetramer, associate with each other in the C1 complex. This analysis also provides the first experimental evidence that, in the proenzyme form of C1, the C1s serine protease domain is partly positioned inside the C1q cone and yields precise information about its orientation in the complex. These results provide further structural insights into the architecture of the C1 complex, allowing significant improvement of our current C1 model.
Figures


Similar articles
-
Identification of the C1q-binding Sites of Human C1r and C1s: a refined three-dimensional model of the C1 complex of complement.J Biol Chem. 2009 Jul 17;284(29):19340-8. doi: 10.1074/jbc.M109.004473. Epub 2009 May 27. J Biol Chem. 2009. PMID: 19473974 Free PMC article.
-
Chemical characterization and location of ionic interactions involved in the assembly of the C1 complex of human complement.J Protein Chem. 1993 Dec;12(6):771-81. doi: 10.1007/BF01024936. J Protein Chem. 1993. PMID: 8136028
-
Expression of recombinant human complement C1q allows identification of the C1r/C1s-binding sites.Proc Natl Acad Sci U S A. 2013 May 21;110(21):8650-5. doi: 10.1073/pnas.1304894110. Epub 2013 May 6. Proc Natl Acad Sci U S A. 2013. PMID: 23650384 Free PMC article.
-
Assembly of the C1 complex.Behring Inst Mitt. 1993 Dec;(93):189-95. Behring Inst Mitt. 1993. PMID: 8172567 Review.
-
C1 subcomponent complexes: basic and clinical aspects.Behring Inst Mitt. 1993 Dec;(93):292-8. Behring Inst Mitt. 1993. PMID: 8172579 Review.
Cited by
-
The EPICC Family of Anti-Inflammatory Peptides: Next Generation Peptides, Additional Mechanisms of Action, and In Vivo and Ex Vivo Efficacy.Front Immunol. 2022 Feb 9;13:752315. doi: 10.3389/fimmu.2022.752315. eCollection 2022. Front Immunol. 2022. PMID: 35222367 Free PMC article. Review.
-
Deciphering the fine details of c1 assembly and activation mechanisms: "mission impossible"?Front Immunol. 2014 Nov 6;5:565. doi: 10.3389/fimmu.2014.00565. eCollection 2014. Front Immunol. 2014. PMID: 25414705 Free PMC article. Review.
-
Complement activation, regulation, and molecular basis for complement-related diseases.EMBO J. 2015 Nov 12;34(22):2735-57. doi: 10.15252/embj.201591881. Epub 2015 Oct 21. EMBO J. 2015. PMID: 26489954 Free PMC article. Review.
-
Procollagen C-endopeptidase Enhancer Protein 2 (PCPE2) Reduces Atherosclerosis in Mice by Enhancing Scavenger Receptor Class B1 (SR-BI)-mediated High-density Lipoprotein (HDL)-Cholesteryl Ester Uptake.J Biol Chem. 2015 Jun 19;290(25):15496-15511. doi: 10.1074/jbc.M115.646240. Epub 2015 May 6. J Biol Chem. 2015. PMID: 25947382 Free PMC article.
-
Reply to Arlaud et al.: Structure of the C1 complex and the unbound C1r2s2 tetramer.Proc Natl Acad Sci U S A. 2017 Jul 18;114(29):E5768-E5770. doi: 10.1073/pnas.1704353114. Epub 2017 Jul 12. Proc Natl Acad Sci U S A. 2017. PMID: 28701383 Free PMC article. No abstract available.
References
-
- Duncan R. C., Wijeyewickrema L. C., Pike R. N. (2008) Biochimie 90, 387–395 - PubMed
-
- Frank M. M., Fries L. F. (1991) Immunol. Today 12, 322–326 - PubMed
-
- Arlaud G. J., Gaboriaud C., Thielens N. M., Rossi V. (2002) Biochem. Soc. Trans. 30, 1001–1006 - PubMed
-
- Arlaud G. J., Thielens N. (1993) Methods Enzymol. 223, 61–82 - PubMed
-
- Gaboriaud C., Teillet F., Gregory L. A., Thielens N. M., Arlaud G. J. (2007) Immunobiology 212, 279–288 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

