The architecture and chemical stability of the archaeal Sulfolobus turreted icosahedral virus
- PMID: 20592081
- PMCID: PMC2937605
- DOI: 10.1128/JVI.00708-10
The architecture and chemical stability of the archaeal Sulfolobus turreted icosahedral virus
Abstract
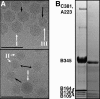
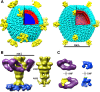
Viruses utilize a diverse array of mechanisms to deliver their genomes into hosts. While great strides have been made in understanding the genome delivery of eukaryotic and prokaryotic viruses, little is known about archaeal virus genome delivery and the associated particle changes. The Sulfolobus turreted icosahedral virus (STIV) is a double-stranded DNA (dsDNA) archaeal virus that contains a host-derived membrane sandwiched between the genome and the proteinaceous capsid shell. Using cryo-electron microscopy (cryo-EM) and different biochemical treatments, we identified three viral morphologies that may correspond to biochemical disassembly states of STIV. One of these morphologies was subtly different from the previously published 27-A-resolution electron density that was interpreted with the crystal structure of the major capsid protein (MCP). However, these particles could be analyzed at 12.5-A resolution by cryo-EM. Comparing these two structures, we identified the location of multiple proteins forming the large turret-like appendages at the icosahedral vertices, observed heterogeneous glycosylation of the capsid shell, and identified mobile MCP C-terminal arms responsible for tethering and releasing the underlying viral membrane to and from the capsid shell. Collectively, our studies allow us to propose a fusogenic mechanism of genome delivery by STIV, in which the dismantled capsid shell allows for the fusion of the viral and host membranes and the internalization of the viral genome.
Figures



Similar articles
-
Maximizing the potential of electron cryomicroscopy data collected using direct detectors.J Struct Biol. 2013 Nov;184(2):193-202. doi: 10.1016/j.jsb.2013.09.003. Epub 2013 Sep 12. J Struct Biol. 2013. PMID: 24036281 Free PMC article.
-
Atomic structure of the 75 MDa extremophile Sulfolobus turreted icosahedral virus determined by CryoEM and X-ray crystallography.Proc Natl Acad Sci U S A. 2013 Apr 2;110(14):5504-9. doi: 10.1073/pnas.1300601110. Epub 2013 Mar 21. Proc Natl Acad Sci U S A. 2013. PMID: 23520050 Free PMC article.
-
Characterization of the archaeal thermophile Sulfolobus turreted icosahedral virus validates an evolutionary link among double-stranded DNA viruses from all domains of life.J Virol. 2006 Aug;80(15):7625-35. doi: 10.1128/JVI.00522-06. J Virol. 2006. PMID: 16840341 Free PMC article.
-
Structure and cell biology of archaeal virus STIV.Curr Opin Virol. 2012 Apr;2(2):122-7. doi: 10.1016/j.coviro.2012.01.007. Epub 2012 Mar 20. Curr Opin Virol. 2012. PMID: 22482708 Free PMC article. Review.
-
Exceptional virion release mechanism: one more surprise from archaeal viruses.Curr Opin Microbiol. 2011 Jun;14(3):315-20. doi: 10.1016/j.mib.2011.04.006. Epub 2011 Apr 30. Curr Opin Microbiol. 2011. PMID: 21531608 Review.
Cited by
-
Maximizing the potential of electron cryomicroscopy data collected using direct detectors.J Struct Biol. 2013 Nov;184(2):193-202. doi: 10.1016/j.jsb.2013.09.003. Epub 2013 Sep 12. J Struct Biol. 2013. PMID: 24036281 Free PMC article.
-
Viral life cycles captured in three-dimensions with electron microscopy tomography.Curr Opin Virol. 2011 Aug;1(2):125-33. doi: 10.1016/j.coviro.2011.06.008. Curr Opin Virol. 2011. PMID: 21887207 Free PMC article. Review.
-
Sulfolobus turreted icosahedral virus c92 protein responsible for the formation of pyramid-like cellular lysis structures.J Virol. 2011 Jul;85(13):6287-92. doi: 10.1128/JVI.00379-11. Epub 2011 Apr 27. J Virol. 2011. PMID: 21525343 Free PMC article.
-
Viruses of the Turriviridae: an emerging model system for studying archaeal virus-host interactions.Front Microbiol. 2023 Sep 21;14:1258997. doi: 10.3389/fmicb.2023.1258997. eCollection 2023. Front Microbiol. 2023. PMID: 37808280 Free PMC article. Review.
-
Structures of enveloped virions determined by cryogenic electron microscopy and tomography.Adv Virus Res. 2019;105:35-71. doi: 10.1016/bs.aivir.2019.07.009. Epub 2019 Aug 20. Adv Virus Res. 2019. PMID: 31522708 Free PMC article.
References
-
- Abrescia, N. G., J. J. Cockburn, J. M. Grimes, G. C. Sutton, J. M. Diprose, S. J. Butcher, S. D. Fuller, C. San Martin, R. M. Burnett, D. I. Stuart, D. H. Bamford, and J. K. Bamford. 2004. Insights into assembly from structural analysis of bacteriophage PRD1. Nature 432:68-74. - PubMed
-
- Abrescia, N. G., J. M. Grimes, H. M. Kivela, R. Assenberg, G. C. Sutton, S. J. Butcher, J. K. Bamford, D. H. Bamford, and D. I. Stuart. 2008. Insights into virus evolution and membrane biogenesis from the structure of the marine lipid-containing bacteriophage PM2. Mol. Cell 31:749-761. - PubMed
-
- Bamford, D. H., J. M. Grimes, and D. I. Stuart. 2005. What does structure tell us about virus evolution? Curr. Opin. Struct. Biol. 15:655-663. - PubMed
-
- Benson, S. D., J. K. H. Bamford, D. H. Bamford, and R. Burnett. 1999. Viral evolution revealed by bacteriophage PRD1 and human adenovirus coat protein structures. Cell 98:825-833. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

