Ribonucleotide reductases of Salmonella typhimurium: transcriptional regulation and differential role in pathogenesis
- PMID: 20593029
- PMCID: PMC2892513
- DOI: 10.1371/journal.pone.0011328
Ribonucleotide reductases of Salmonella typhimurium: transcriptional regulation and differential role in pathogenesis
Abstract
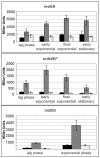
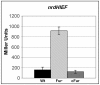
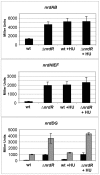
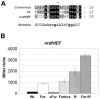
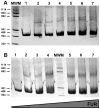
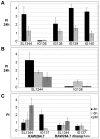
Ribonucleotide reductases (RNRs) are essential enzymes that carry out the de novo synthesis of deoxyribonucleotides by reducing ribonucleotides. There are three different classes of RNRs (I, II and III), all having different oxygen dependency and biochemical characteristics. Salmonella enterica serovar Typhimurium (S. Typhimurium) harbors class Ia, class Ib and class III RNRs in its genome. We have studied the transcriptional regulation of these three RNR classes in S. Typhimurium as well as their differential function during infection of macrophage and epithelial cells. Deletion of both NrdR and Fur, two main transcriptional regulators, indicates that Fur specifically represses the class Ib enzyme and that NrdR acts as a global repressor of all three classes. A Fur recognition sequence within the nrdHIEF promoter has also been described and confirmed by electrophoretic mobility shift assays (EMSA). In order to elucidate the role of each RNR class during infection, S. Typhimurium single and double RNR mutants (as well as Fur and NrdR mutants) were used in infection assays with macrophage and epithelial cell lines. Our results indicate class Ia to be mainly responsible for deoxyribonucleotide production during invasion and proliferation inside macrophages and epithelial cells. Neither class Ib nor class III seem to be essential for growth under these conditions. However, class Ib is able to maintain certain growth in an nrdAB mutant during the first hours of macrophage infection. Our results suggest that, during the early stages of macrophage infection, class Ib may contribute to deoxyribonucleotide synthesis by means of both an NrdR and a Fur-dependent derepression of nrdHIEF due to hydrogen peroxide production and DNA damage associated with the oxidative burst, thus helping to overcome the host defenses.
Conflict of interest statement
Figures


Similar articles
-
Ribonucleotide reductases: essential enzymes for bacterial life.Front Cell Infect Microbiol. 2014 Apr 28;4:52. doi: 10.3389/fcimb.2014.00052. eCollection 2014. Front Cell Infect Microbiol. 2014. PMID: 24809024 Free PMC article. Review.
-
NrdR controls differential expression of the Escherichia coli ribonucleotide reductase genes.J Bacteriol. 2007 Jul;189(14):5012-21. doi: 10.1128/JB.00440-07. Epub 2007 May 11. J Bacteriol. 2007. PMID: 17496099 Free PMC article.
-
Alternative oxygen-dependent and oxygen-independent ribonucleotide reductases in Streptomyces: cross-regulation and physiological role in response to oxygen limitation.Mol Microbiol. 2004 Nov;54(4):1022-35. doi: 10.1111/j.1365-2958.2004.04325.x. Mol Microbiol. 2004. PMID: 15522084
-
The Streptomyces NrdR transcriptional regulator is a Zn ribbon/ATP cone protein that binds to the promoter regions of class Ia and class II ribonucleotide reductase operons.J Bacteriol. 2006 Nov;188(21):7635-44. doi: 10.1128/JB.00903-06. Epub 2006 Sep 1. J Bacteriol. 2006. PMID: 16950922 Free PMC article.
-
Deoxyribonucleotide synthesis in anaerobic microorganisms: the class III ribonucleotide reductase.Prog Nucleic Acid Res Mol Biol. 2002;72:95-127. doi: 10.1016/s0079-6603(02)72068-0. Prog Nucleic Acid Res Mol Biol. 2002. PMID: 12206460 Review.
Cited by
-
Nitric oxide and salmonella pathogenesis.Front Microbiol. 2011 Apr 20;2:84. doi: 10.3389/fmicb.2011.00084. eCollection 2011. Front Microbiol. 2011. PMID: 21833325 Free PMC article.
-
Genome sequence of Thermotoga sp. strain RQ2, a hyperthermophilic bacterium isolated from a geothermally heated region of the seafloor near Ribeira Quente, the Azores.J Bacteriol. 2011 Oct;193(20):5869-70. doi: 10.1128/JB.05923-11. J Bacteriol. 2011. PMID: 21952543 Free PMC article.
-
CT406 encodes a chlamydial ortholog of NrdR, a repressor of ribonucleotide reductase.J Bacteriol. 2011 Sep;193(17):4396-404. doi: 10.1128/JB.00294-11. Epub 2011 Jul 1. J Bacteriol. 2011. PMID: 21725017 Free PMC article.
-
Ribonucleotide reductases: essential enzymes for bacterial life.Front Cell Infect Microbiol. 2014 Apr 28;4:52. doi: 10.3389/fcimb.2014.00052. eCollection 2014. Front Cell Infect Microbiol. 2014. PMID: 24809024 Free PMC article. Review.
-
Escherichia coli class Ib ribonucleotide reductase contains a dimanganese(III)-tyrosyl radical cofactor in vivo.Biochemistry. 2011 Mar 15;50(10):1672-81. doi: 10.1021/bi101881d. Epub 2011 Feb 15. Biochemistry. 2011. PMID: 21250660 Free PMC article.
References
-
- Jordan A, Reichard P. Ribonucleotide reductases. Annu Rev Biochem. 1998;67:71–98. - PubMed
-
- Nordlund P, Reichard P. Ribonucleotide reductases. Annu Rev Biochem. 2006;75:681–706. - PubMed
-
- Jacobson BA, Fuchs JA. A 45 bp inverted repeat is required for cell cycle regulation of the Escherichia coli nrd operon. Mol Microbiol. 1998;28:1307–1314. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

