Characterization of the six glycosyltransferases involved in the biosynthesis of Yersinia enterocolitica serotype O:3 lipopolysaccharide outer core
- PMID: 20595390
- PMCID: PMC2934697
- DOI: 10.1074/jbc.M110.111336
Characterization of the six glycosyltransferases involved in the biosynthesis of Yersinia enterocolitica serotype O:3 lipopolysaccharide outer core
Abstract
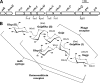
Yersinia enterocolitica (Ye) is a gram-negative bacterium; Ye serotype O:3 expresses lipopolysaccharide (LPS) with a hexasaccharide branch known as the outer core (OC). The OC is important for the resistance of the bacterium to cationic antimicrobial peptides and also functions as a receptor for bacteriophage phiR1-37 and enterocoliticin. The biosynthesis of the OC hexasaccharide is directed by the OC gene cluster that contains nine genes (wzx, wbcKLMNOPQ, and gne). In this study, we inactivated the six OC genes predicted to encode glycosyltransferases (GTase) one by one by nonpolar mutations to assign functions to their gene products. The mutants expressed no OC or truncated OC oligosaccharides of different lengths. The truncated OC oligosaccharides revealed that the minimum structural requirements for the interactions of OC with bacteriophage phiR1-37, enterocoliticin, and OC-specific monoclonal antibody 2B5 were different. Furthermore, using chemical and structural analyses of the mutant LPSs, we could assign specific functions to all six GTases and also revealed the exact order in which the transferases build the hexasaccharide. Comparative modeling of the catalytic sites of glucosyltransferases WbcK and WbcL followed by site-directed mutagenesis allowed us to identify Asp-182 and Glu-181, respectively, as catalytic base residues of these two GTases. In general, conclusive evidence for specific GTase functions have been rare due to difficulties in accessibility of the appropriate donors and acceptors; however, in this work we were able to utilize the structural analysis of LPS to get direct experimental evidence for five different GTase specificities.
Figures

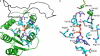
Similar articles
-
Functional characterization of Gne (UDP-N-acetylglucosamine-4-epimerase), Wzz (chain length determinant), and Wzy (O-antigen polymerase) of Yersinia enterocolitica serotype O:8.J Bacteriol. 2002 Aug;184(15):4277-87. doi: 10.1128/JB.184.15.4277-4287.2002. J Bacteriol. 2002. PMID: 12107146 Free PMC article.
-
Characterization and biological role of the O-polysaccharide gene cluster of Yersinia enterocolitica serotype O:9.J Bacteriol. 2007 Oct;189(20):7244-53. doi: 10.1128/JB.00605-07. Epub 2007 Aug 10. J Bacteriol. 2007. PMID: 17693522 Free PMC article.
-
Mutations in the genes for synthesis of the outer core region of the lipopolysaccharide of Yersinia enterocolitica O:3.J Appl Microbiol. 2003;94(4):686-92. doi: 10.1046/j.1365-2672.2003.01897.x. J Appl Microbiol. 2003. PMID: 12631204
-
The biosynthesis and biological role of lipopolysaccharide O-antigens of pathogenic Yersiniae.Carbohydr Res. 2003 Nov 14;338(23):2521-9. doi: 10.1016/s0008-6215(03)00305-7. Carbohydr Res. 2003. PMID: 14670713 Review.
-
Molecular genetics and biochemistry of Yersinia lipopolysaccharide.APMIS. 1996 Dec;104(12):849-72. APMIS. 1996. PMID: 9048864 Review.
Cited by
-
φYeO3-12 phage tail fiber Gp17 as a promising high specific tool for recognition of Yersinia enterocolitica pathogenic serotype O:3.AMB Express. 2022 Jan 6;12(1):1. doi: 10.1186/s13568-021-01341-2. AMB Express. 2022. PMID: 34989907 Free PMC article.
-
Identification of the lipopolysaccharide core of Yersinia pestis and Yersinia pseudotuberculosis as the receptor for bacteriophage φA1122.J Bacteriol. 2011 Sep;193(18):4963-72. doi: 10.1128/JB.00339-11. Epub 2011 Jul 15. J Bacteriol. 2011. PMID: 21764935 Free PMC article.
-
The Podovirus ϕ80-18 Targets the Pathogenic American Biotype 1B Strains of Yersinia enterocolitica.Front Microbiol. 2020 Jun 19;11:1356. doi: 10.3389/fmicb.2020.01356. eCollection 2020. Front Microbiol. 2020. PMID: 32636826 Free PMC article.
-
Identification and characterization of six glycosyltransferases involved in the biosynthesis of a new bacterial exopolysaccharide in Paenibacillus elgii.Appl Microbiol Biotechnol. 2018 Feb;102(3):1357-1366. doi: 10.1007/s00253-017-8673-y. Epub 2017 Dec 3. Appl Microbiol Biotechnol. 2018. PMID: 29199353
-
RNA-Sequencing Reveals the Progression of Phage-Host Interactions between φR1-37 and Yersinia enterocolitica.Viruses. 2016 Apr 22;8(4):111. doi: 10.3390/v8040111. Viruses. 2016. PMID: 27110815 Free PMC article.
References
-
- Holst O., Müller-Loennies S. (2007) in Comprehensive Glycoscience: From Chemistry to Systems Biology (Kamerling J. P., Boons G.-J., Lee Y. C., Suzuki A., Taniguchi N., Voragen A. G. J. eds) Vol. 1, pp. 123–175, Elsevier, Oxford, UK
-
- Skurnik M., Venho R., Bengoechea J. A., Moriyón I. (1999) Mol. Microbiol. 31, 1443–1462 - PubMed
-
- Pinta E., Duda K. A., Hanuszkiewicz A., Kaczyński Z., Lindner B., Miller W. L., Hyytiäinen H., Vogel C., Borowski S., Kasperkiewicz K., Lam J. S., Radziejewska-Lebrecht J., Skurnik M., Holst O. (2009) Chemistry 15, 9747–9754 - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

