Selective detection of Cathepsin E proteolytic activity
- PMID: 20600629
- PMCID: PMC2914798
- DOI: 10.1016/j.bbagen.2010.06.005
Selective detection of Cathepsin E proteolytic activity
Abstract
Background: Aspartic proteases Cathepsin (Cath) E and D are two different proteases, but they share many common characteristics, including molecular weight, catalytic mechanism, substrate preferences, proteolytic conditions and inhibition susceptibility. To define the biological roles of these proteases, it is necessary to elucidate their substrate specificity. In the present study, we report a new peptide-substrate that is only sensitive to Cath E but not Cath D.
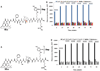
Methods: Substrate e, Mca-Ala-Gly-Phe-Ser-Leu-Pro-Ala-Lys(Dnp)-DArg-CONH₂, designed in such a way that due to the close proximity of a Mca-donor and a Dnp-acceptor, near complete intramolecular quenching effect was achieved in its intact state. After the proteolytic cleavage of the hydrophobic motif of peptide substrate, both Mca and Dnp would be further apart, resulting in bright fluorescence.
Results: Substrate e showed a 265 fold difference in the net fluorescence signals between Cath E and D. This Cath E selectivity was established by having -Leu**Pro- residues at the scissile peptide bond. The confined cleavage site of substrate e was confirmed by LC-MS. The catalytic efficiency (K(cat)/K(M)) of Cath E for substrate e was 16.7 μM⁻¹S⁻¹. No measurable catalytic efficiency was observed using Cath D and no detectable fluorescent changes when incubated with Cath S and Cath B.
Conclusions: This study demonstrated the promise of using the developed fluorogenic substrate e as a selective probe for Cath E proteolytic activity measurement.
General significance: This study forms the foundation of Cath E specific inhibitor development in further studies.
Copyright © 2010 Elsevier B.V. All rights reserved.
Figures
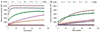




Similar articles
-
Characterization of new fluorogenic substrates for the rapid and sensitive assay of cathepsin E and cathepsin D.J Biochem. 1999 Jun;125(6):1137-43. doi: 10.1093/oxfordjournals.jbchem.a022396. J Biochem. 1999. PMID: 10348917
-
Design, synthesis, and bioactivities of tasiamide B derivatives as cathepsin D inhibitors.J Pept Sci. 2019 Apr;25(4):e3154. doi: 10.1002/psc.3154. Epub 2019 Feb 7. J Pept Sci. 2019. PMID: 30734395
-
[p-Nitroanilides of amino acids and peptides and fluorescence peptide with inner fluorescence quenching as substrates for cathepsins H, B, D and high molecular weight aspartic peptidase in the brain].Biokhimiia. 1987 Dec;52(12):2033-7. Biokhimiia. 1987. PMID: 3328984 Russian.
-
Pathophysiological functions of cathepsin D: Targeting its catalytic activity versus its protein binding activity?Biochimie. 2010 Nov;92(11):1635-43. doi: 10.1016/j.biochi.2010.05.009. Epub 2010 May 21. Biochimie. 2010. PMID: 20493920 Review.
-
Cathepsin H: Molecular characteristics and clues to function and mechanism.Biochem Pharmacol. 2023 Jun;212:115585. doi: 10.1016/j.bcp.2023.115585. Epub 2023 May 5. Biochem Pharmacol. 2023. PMID: 37148981 Review.
Cited by
-
Recombinant protein susceptibility to proteolysis in the plant cell secretory pathway is pH-dependent.Plant Biotechnol J. 2018 Nov;16(11):1928-1938. doi: 10.1111/pbi.12928. Epub 2018 May 2. Plant Biotechnol J. 2018. PMID: 29618167 Free PMC article.
-
Peptide aptamer-modified single-walled carbon nanotube-based transistors for high-performance biosensors.Sci Rep. 2017 Dec 20;7(1):17881. doi: 10.1038/s41598-017-18169-1. Sci Rep. 2017. PMID: 29263412 Free PMC article.
-
Biochemical characterization and structural modeling of human cathepsin E variant 2 in comparison to the wild-type protein.Biol Chem. 2012 Mar;393(3):177-86. doi: 10.1515/hsz-2011-0219. Biol Chem. 2012. PMID: 22718633 Free PMC article.
-
Pancreatic cancer-associated Cathepsin E as a drug activator.J Control Release. 2013 May 10;167(3):221-7. doi: 10.1016/j.jconrel.2013.02.007. Epub 2013 Feb 26. J Control Release. 2013. PMID: 23422726 Free PMC article.
-
Designing and developing S100P inhibitor 5-methyl cromolyn for pancreatic cancer therapy.Mol Cancer Ther. 2013 May;12(5):654-62. doi: 10.1158/1535-7163.MCT-12-0771. Epub 2013 Jan 9. Mol Cancer Ther. 2013. PMID: 23303403 Free PMC article.
References
-
- Foltmann B, Szecsi PB. In: Handbook of Proteolytic Enzymes. Barrett AJ, Rawlings ND, Woessner JF, editors. San Diego: Academic Press; 1998. pp. 819–823.
-
- Tang J. In: Handbook of Proteolytic Enzymes. Barrett AJ, Rawlings ND, Woessner JF, editors. San Diego: Academic Press; 1998. pp. 828–836.
-
- Kageyama T, Takahashi K. J Biochem. 1980;87:725–735. - PubMed
-
- Muto N, Arai KM, Tani S. Biochim Biophys Acta. 1983;745:61–69. - PubMed
-
- Bennett K, Levine T, Ellis JS, Peanasky RJ, Samloff IM, Kay J, Chain BM. Eur J Immunol. 1992;22:1519–1524. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

