Fast, high-contrast imaging of animal development with scanned light sheet-based structured-illumination microscopy
- PMID: 20601950
- PMCID: PMC4418465
- DOI: 10.1038/nmeth.1476
Fast, high-contrast imaging of animal development with scanned light sheet-based structured-illumination microscopy
Abstract
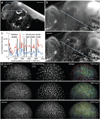
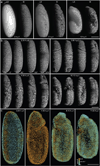
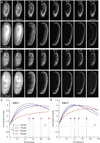
Recording light-microscopy images of large, nontransparent specimens, such as developing multicellular organisms, is complicated by decreased contrast resulting from light scattering. Early zebrafish development can be captured by standard light-sheet microscopy, but new imaging strategies are required to obtain high-quality data of late development or of less transparent organisms. We combined digital scanned laser light-sheet fluorescence microscopy with incoherent structured-illumination microscopy (DSLM-SI) and created structured-illumination patterns with continuously adjustable frequencies. Our method discriminates the specimen-related scattered background from signal fluorescence, thereby removing out-of-focus light and optimizing the contrast of in-focus structures. DSLM-SI provides rapid control of the illumination pattern, exceptional imaging quality and high imaging speeds. We performed long-term imaging of zebrafish development for 58 h and fast multiple-view imaging of early Drosophila melanogaster development. We reconstructed cell positions over time from the Drosophila DSLM-SI data and created a fly digital embryo.
Figures

References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

