ARRDC3 suppresses breast cancer progression by negatively regulating integrin beta4
- PMID: 20603614
- PMCID: PMC2997682
- DOI: 10.1038/onc.2010.250
ARRDC3 suppresses breast cancer progression by negatively regulating integrin beta4
Abstract
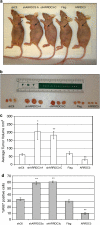
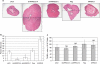
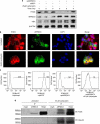
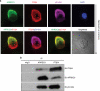
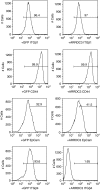
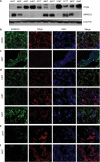
Large-scale genetic analyses of human tumor samples have been used to identify novel oncogenes, tumor suppressors and prognostic factors, but the functions and molecular interactions of many individual genes have not been determined. In this study we examined the cellular effects and molecular mechanism of the arrestin family member, ARRDC3, a gene preferentially lost in a subset of breast cancers. Oncomine data revealed that the expression of ARRDC3 decreases with tumor grade, metastases and recurrences. ARRDC3 overexpression represses cancer cell proliferation, migration, invasion, growth in soft agar and in vivo tumorigenicity, whereas downregulation of ARRCD3 has the opposite effects. Mechanistic studies showed that ARRDC3 functions in a novel regulatory pathway that controls the cell surface adhesion molecule, beta-4 integrin (ITGbeta4), a protein associated with aggressive tumor behavior. Our data indicates ARRDC3 directly binds to a phosphorylated form of ITGbeta4 leading to its internalization, ubiquitination and ultimate degradation. The results identify the ARRCD3-ITGbeta4 pathway as a new therapeutic target in breast cancer and show the importance of connecting genetic arrays with mechanistic studies in the search for new treatments.
Figures



Similar articles
-
ARRDC3 Inhibits the Progression of Human Prostate Cancer Through ARRDC3-ITGβ4 Pathway.Curr Mol Med. 2017;17(3):221-229. doi: 10.2174/1566524017666170807144711. Curr Mol Med. 2017. PMID: 28782483
-
miR-221/222 control luminal breast cancer tumor progression by regulating different targets.Cell Cycle. 2014;13(11):1811-26. doi: 10.4161/cc.28758. Epub 2014 Apr 15. Cell Cycle. 2014. PMID: 24736554 Free PMC article.
-
The α-arrestin ARRDC3 suppresses breast carcinoma invasion by regulating G protein-coupled receptor lysosomal sorting and signaling.J Biol Chem. 2018 Mar 2;293(9):3350-3362. doi: 10.1074/jbc.RA117.001516. Epub 2018 Jan 18. J Biol Chem. 2018. PMID: 29348172 Free PMC article.
-
The α-Arrestin ARRDC3 Is an Emerging Multifunctional Adaptor Protein in Cancer.Antioxid Redox Signal. 2022 May;36(13-15):1066-1079. doi: 10.1089/ars.2021.0193. Epub 2022 Jan 4. Antioxid Redox Signal. 2022. PMID: 34465145 Free PMC article. Review.
-
[Molecular mechanisms of breast cancer progression].Nihon Geka Gakkai Zasshi. 1996 May;97(5):381-6. Nihon Geka Gakkai Zasshi. 1996. PMID: 8709941 Review. Japanese.
Cited by
-
α-Arrestin ARRDC3 tumor suppressor function is linked to GPCR-induced TAZ activation and breast cancer metastasis.J Cell Sci. 2021 Apr 15;134(8):jcs254888. doi: 10.1242/jcs.254888. Epub 2021 Apr 22. J Cell Sci. 2021. PMID: 33722977 Free PMC article.
-
Molecular Biomarkers for Contemporary Therapies in Hormone Receptor-Positive Breast Cancer.Genes (Basel). 2021 Feb 17;12(2):285. doi: 10.3390/genes12020285. Genes (Basel). 2021. PMID: 33671468 Free PMC article. Review.
-
TransCONFIRM: Identification of a Genetic Signature of Response to Fulvestrant in Advanced Hormone Receptor-Positive Breast Cancer.Clin Cancer Res. 2016 Dec 1;22(23):5755-5764. doi: 10.1158/1078-0432.CCR-16-0148. Epub 2016 May 16. Clin Cancer Res. 2016. PMID: 27185372 Free PMC article. Clinical Trial.
-
Inhibition of Integrin-HER2 signaling by Cucurbitacin B leads to in vitro and in vivo breast tumor growth suppression.Oncotarget. 2014 Apr 15;5(7):1812-28. doi: 10.18632/oncotarget.1743. Oncotarget. 2014. PMID: 24729020 Free PMC article.
-
Integrins and Cell Metabolism: An Intimate Relationship Impacting Cancer.Int J Mol Sci. 2017 Jan 18;18(1):189. doi: 10.3390/ijms18010189. Int J Mol Sci. 2017. PMID: 28106780 Free PMC article. Review.
References
-
- Adelaide J, Finetti P, Bekhouche I, Repellini L, Geneix J, Sircoulomb F, et al. Integrated profiling of basal and luminal breast cancers. Cancer Res. 2007;67:11565–11575. - PubMed
-
- Arnaout MA, Mahalingam B, Xiong JP. Integrin structure, allostery, and bidirectional signaling. Annu Rev Cell Dev Biol. 2005;21:381–410. - PubMed
-
- Baril P, Gangeswaran R, Mahon PC, Caulee K, Kocher HM, Harada T, et al. Periostin promotes invasiveness and resistance of pancreatic cancer cells to hypoxia-induced cell death: role of the beta4 integrin and the PI3k pathway. Oncogene. 2007;26:2082–2094. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

