Counter-selection recombineering of the baculovirus genome: a strategy for seamless modification of repeat-containing BACs
- PMID: 20621982
- PMCID: PMC2938205
- DOI: 10.1093/nar/gkq596
Counter-selection recombineering of the baculovirus genome: a strategy for seamless modification of repeat-containing BACs
Abstract
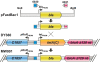
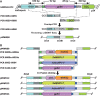
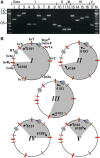
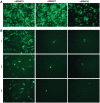
Recombineering is employed to modify large DNA clones such as fosmids, BACs and PACs. Subtle and seamless modifications can be achieved using counter-selection strategies in which a donor cassette carrying both positive and negative markers inserted in the target clone is replaced by the desired sequence change. We are applying counter-selection recombineering to modify bacmid bMON14272, a recombinant baculoviral genome, as we wish to engineer the virus into a therapeutically useful gene delivery vector with cell targeting characteristics. Initial attempts to replace gp64 with Fusion (F) genes from other baculoviruses resulted in many rearranged clones in which the counter-selection cassette had been deleted. Bacmid bMON14272 contains nine highly homologous regions (hrs) and deletions were mapped to recombination between hr pairs. Recombineering modifications were attempted to decrease intramolecular recombination and/or increase recombineering efficiency. Of these only the use of longer homology arms on the donor molecule proved effective permitting seamless modification. bMON14272, because of the presence of the hr sequences, can be considered equivalent to a highly repetitive BAC and, as such, the optimized method detailed here should prove useful to others applying counter-selection recombineering to modify BACs or PACs containing similar regions of significant repeating homologies.
Figures
Similar articles
-
Scarless Baculovirus Genome Editing Using Lambda-Red Recombineering in E. coli.Methods Mol Biol. 2024;2829:109-126. doi: 10.1007/978-1-0716-3961-0_8. Methods Mol Biol. 2024. PMID: 38951330
-
A simplified counter-selection recombineering protocol for creating fluorescent protein reporter constructs directly from C. elegans fosmid genomic clones.BMC Biotechnol. 2013 Jan 3;13:1. doi: 10.1186/1472-6750-13-1. BMC Biotechnol. 2013. PMID: 23281894 Free PMC article.
-
Transgene Recombineering in Bacterial Artificial Chromosomes.Methods Mol Biol. 2019;1874:43-69. doi: 10.1007/978-1-4939-8831-0_3. Methods Mol Biol. 2019. PMID: 30353507
-
Bacterial artificial chromosome mutagenesis using recombineering.J Biomed Biotechnol. 2011;2011:971296. doi: 10.1155/2011/971296. Epub 2010 Dec 9. J Biomed Biotechnol. 2011. PMID: 21197472 Free PMC article. Review.
-
Recombineering: genetic engineering in bacteria using homologous recombination.Curr Protoc Mol Biol. 2014 Apr 14;106:1.16.1-1.16.39. doi: 10.1002/0471142727.mb0116s106. Curr Protoc Mol Biol. 2014. PMID: 24733238 Review.
Cited by
-
High-efficiency counterselection recombineering for site-directed mutagenesis in bacterial artificial chromosomes.Nat Methods. 2011 Dec 4;9(1):103-9. doi: 10.1038/nmeth.1803. Nat Methods. 2011. PMID: 22138824
-
Improved seamless mutagenesis by recombineering using ccdB for counterselection.Nucleic Acids Res. 2014 Mar;42(5):e37. doi: 10.1093/nar/gkt1339. Epub 2013 Dec 24. Nucleic Acids Res. 2014. PMID: 24369425 Free PMC article.
-
Scarless Baculovirus Genome Editing Using Lambda-Red Recombineering in E. coli.Methods Mol Biol. 2024;2829:109-126. doi: 10.1007/978-1-0716-3961-0_8. Methods Mol Biol. 2024. PMID: 38951330
-
Seamless replacement of Autographa californica multiple nucleopolyhedrovirus gp64 with each of five novel type II alphabaculovirus fusion sequences generates pseudotyped virus that fails to transduce mammalian cells.J Gen Virol. 2012 Jul;93(Pt 7):1583-1590. doi: 10.1099/vir.0.041921-0. Epub 2012 Apr 4. J Gen Virol. 2012. PMID: 22492915 Free PMC article.
-
Lambda red mediated gap repair utilizes a novel replicative intermediate in Escherichia coli.PLoS One. 2015 Mar 24;10(3):e0120681. doi: 10.1371/journal.pone.0120681. eCollection 2015. PLoS One. 2015. PMID: 25803509 Free PMC article.
References
-
- Court DL, Sawitzke JA, Thomason LC. Genetic engineering using homologous recombination. Annu. Rev. Genet. 2002;36:361–388. - PubMed
-
- Zhang Y, Buchholz F, Muyrers JP, Stewart AF. A new logic for DNA engineering using recombination in Escherichia coli. Nat. Genet. 1998;20:123–128. - PubMed
-
- Copeland NG, Jenkins NA, Court DL. Recombineering: a powerful new tool for mouse functional genomics. Nat. Rev. Genet. 2001;2:769–779. - PubMed
-
- Lee EC, Yu D, Martinez de Velasco J, Tessarollo L, Swing DA, Court DL, Jenkins NA, Copeland NG. A highly efficient Escherichia coli-based chromosome engineering system adapted for recombinogenic targeting and subcloning of BAC DNA. Genomics. 2001;73:56–65. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

