Systems-level metabolic flux profiling elucidates a complete, bifurcated tricarboxylic acid cycle in Clostridium acetobutylicum
- PMID: 20622067
- PMCID: PMC2937365
- DOI: 10.1128/JB.00490-10
Systems-level metabolic flux profiling elucidates a complete, bifurcated tricarboxylic acid cycle in Clostridium acetobutylicum
Erratum in
- J Bacteriol. 2011 Dec;193(23):6805
Abstract
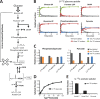
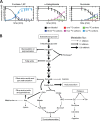
Obligatory anaerobic bacteria are major contributors to the overall metabolism of soil and the human gut. The metabolic pathways of these bacteria remain, however, poorly understood. Using isotope tracers, mass spectrometry, and quantitative flux modeling, here we directly map the metabolic pathways of Clostridium acetobutylicum, a soil bacterium whose major fermentation products include the biofuels butanol and hydrogen. While genome annotation suggests the absence of most tricarboxylic acid (TCA) cycle enzymes, our results demonstrate that this bacterium has a complete, albeit bifurcated, TCA cycle; oxaloacetate flows to succinate both through citrate/alpha-ketoglutarate and via malate/fumarate. Our investigations also yielded insights into the pathways utilized for glucose catabolism and amino acid biosynthesis and revealed that the organism's one-carbon metabolism is distinct from that of model microbes, involving reversible pyruvate decarboxylation and the use of pyruvate as the one-carbon donor for biosynthetic reactions. This study represents the first in vivo characterization of the TCA cycle and central metabolism of C. acetobutylicum. Our results establish a role for the full TCA cycle in an obligatory anaerobic organism and demonstrate the importance of complementing genome annotation with isotope tracer studies for determining the metabolic pathways of diverse microbes.
Figures


References
-
- Bajad, S. U., W. Lu, E. H. Kimball, J. Yuan, C. Peterson, and J. D. Rabinowitz. 2006. Separation and quantitation of water soluble cellular metabolites by hydrophilic interaction chromatography-tandem mass spectrometry. J. Chromatogr. A 1125:76-88. - PubMed
-
- Brinkac, L. M., T. Davidsen, E. Beck, A. Ganapathy, E. Caler, R. J. Dodson, A. S. Durkin, D. M. Harkins, H. Lorenzi, R. Madupu, Y. Sebastian, S. Shrivastava, M. Thiagarajan, J. Orvis, J. P. Sundaram, J. Crabtree, K. Galens, Y. Zhao, J. M. Inman, R. Montgomery, S. Schobel, K. Galinsky, D. M. Tanenbaum, A. Resnick, N. Zafar, O. White, and G. Sutton. 2010. Pathema: a clade-specific bioinformatics resource center for pathogen research. Nucleic Acids Res. 38:D408-D414. - PMC - PubMed
-
- Dehal, P. S., M. P. Joachimiak, M. N. Price, J. T. Bates, J. K. Baumohl, D. Chivian, G. D. Friedland, K. H. Huang, K. Keller, P. S. Novichkov, I. L. Dubchak, E. J. Alm, and A. P. Arkin. 2010. MicrobesOnline: an integrated portal for comparative and functional genomics. Nucleic Acids Res. 38:D396-D400. - PMC - PubMed
-
- Dürre, P. 2005. Handbook on clostridia. Taylor & Francis, Boca Raton, FL.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

